search this blog
Friday, March 13, 2015
Yamnaya-related ancestry proportions in Europe and west Asia
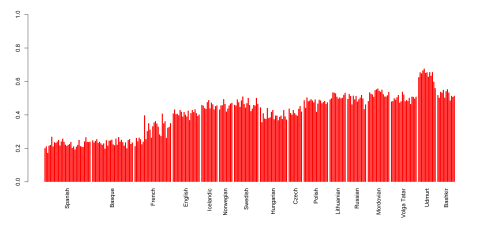
Here's a quick and dirty attempt to flush out a Yamnaya-specific ancestral component with the ADMIXTURE software and a few Yamnaya genomes from the recent Haak et al. paper: K6 spreadsheet.
Obviously, we'll need many more ancient samples from the vast Yamnaya horizon to be able to estimate direct Yamnaya ancestry in modern populations with any great confidence. But I'd say this looks like a very reasonable attempt, with more or less comparable results to those published by Haak et al. (for instance, see Figure 3 from the study here).
Please note that this wasn't a supervised run. In other words, I didn't mark the Yamnaya genomes as reference samples with the aim of creating a cluster from them.
However, I initially excluded all individuals from northeastern Europe, the north Caucasus and South Asia from the analysis. The reason I did this was because samples from these regions have a peculiar habit of creating very robust clusters in ADMIXTURE, which is useful when looking at recent variation and wanting low cross validation errors, but not so great when trying to resurrect genetic components from the depths of prehistory.
Once I had a dataset that was forcing the algorithm to focus its attention on the ancient genomes and producing consistent results, I tested the problem samples in batches of 5-10, thus making sure they didn't skew the analysis.
Interestingly, the Yamnaya-specific component peaks in Udmurts, who live close to where the Yamnaya samples were collected. This can hardly be a coincidence.
In any case, I'm hoping to look at this issue in more detail soon with the help of qpAdm, a new program released recently with the updated ADMIXTOOLS package (see here). Based on f4 statistics, qpAdm is specifically designed for analyzing ancient admixture events.
Citation...
Haak et al., Massive migration from the steppe was a source for Indo-European languages in Europe, Nature, Advance online publication, doi:10.1038/nature14317
Subscribe to:
Post Comments (Atom)

240 comments:
1 – 200 of 240 Newer› Newest»David
Well done on yr effort
Are these result based on a comparison which "sets" Yamnaya as ancestral and then seeks proportions or is it done through independent analysis of all the data ?
Thanks David. I'm just looking at it now, but a quick question about the clusters:
I suppose that pre-Yamnaya means something like European Middle Neolithic, and Middle Eastern is the same as the old ENF to catch the excess of Basal Eurasian, right?
Mike,
It was an unsupervised run. The software picked the clusters.
Alberto,
Middle Eastern makes up about 25% of Stutgart. So I'd say it was the main component in the Neolithic Near East.
On the other hand, I suspect that the Cardial Neolithic groups had more WHG-related ancestry of different types, and this is why Spain_EN comes out mostly pre-Yamnaya here.
The Iceman compared to Stuttgart and Sardinians is almost fully "pre-Yamnaya", but in PCA's and overall affinities it should not differ that much from them. There probably is considerable overlap between pre-Yamnaya and Near Eastern.
Does the "ENA" peak in Oroqen/Hezhen or in Southeast Asia? Given that some Basques and South French have close to 3% and it's so common in Jordanians it seems to be overlapping or mixed as well.
Could you post the Fst-distances?
ENA peaks in the Ami and Atayal. But I don't know what it represents. It's not really important here.
Pre-Yamnaya seems to be a mixture of Near Eastern Neolithic and WHG. But again, I don't think it's all that relevant here.
I'll post the fst matrix next time I run the test.
I guess that sticking to the ancient genomes we do have, this is the best objective approach. Like in the paper, it assumes (since it's not proven otherwise for now) that Yamnaya is the only source of ANE in Europe, so the results are consistent.
Maybe in a couple of weeks that conference about Greece during the LN and BA might give us another data point (and hopefully soon some genomes) that can help us complete the puzzle.
I still think that the K9 experiment (with all the caveats of being an experiment and therefore less objective or scientific) will be closer to the final picture. But for that we'll have to wait and see.
Well, it's a Yamnaya-related or at most Yamnaya-specific component.
So it's not implied here that anyone who has it also has Yamnaya ancestry. Although it sure looks like steppe ancestry from around the southern Urals.
But Yamnaya related could be anything more or less similar in the make up of yamnaya itself . So basically any mixture of EHG-type and "west asian" type . This needn't have come from the steppe ; ultimately or even proximately
In fact ; it probably didn't . Observe that ~ 50% of the late Neolithic- Bronze Age German samples are actually I2 ; 2 are R1a - which could be anywhere from Karelia to the Black Sea ; and only one R1b -(Italo-Celtic branch) from the BB sample .
It's is Yamnaya related, but actually to the non-EHG part of Yamnaya.
The easiest way to see that we are missing an important piece is that Pathans are 35% Yamnaya related, yet they have virtually no WHG or EHG ancestry. So all their Yamnaya comes from the Tajik/Tabassaran part.
The K9 experiment showed how little EHG related ancestry there is everywhere except Siberia. If Yamanya was 50% EHG, it means that the real Yamanaya related ancestry is very low in Europe (ironically, including the Pontic steppe, that must have suffered a big west to east migration at some point) and Central and South Asia.
But again, we need ancient DNA to give credit to this. With what we have now, I can admit that this approach is a valid one as long as it's taken as an ongoing (and far from definitive) process.
Mike,
This is a very peculiar component that is virtually missing from Neolithic Germany, Hungary and even the Copper Age Alps. It's also at overall low levels, and rather choppy levels too, in the Near East today, which implies it got there very late.
So the Eurasian steppe looks like the best source at this point. You can pick your side of the Urals, but that's about as much leeway as there is.
David
I understand what your stats show, but I've stated before - it's an utter impossibility .
So we need to reevaluate what we think we're calculating
See Alberto's statement above
"The K9 experiment showed how little EHG related ancestry there is everywhere except Siberia. If Yamanya was 50% EHG, it means that the real Yamanaya related ancestry is very low in Europe (ironically, including the Pontic steppe, that must have suffered a big west to east migration at some point) and Central and South Asia."
Yamnaya weren't 50% EHG in the K9 experiment. Their EHG-like parts were split between WHG, EHG and Central Asian. So there's no real contradiction and the Yamnaya estimates for Europeans in this test do seem to match pretty well the Haak ones from table S9.27.
If we take EHG to be one parent of Yamnaya, then the other parent would probably be a sort of Central Asian or South Central Asian with the following profile:
ANE 31.06
ASE 5.84
WHG 18.88
ENF 44.22
That's very close to modern South Central Asians. In fact, it's one admixture step away from certain North Indian populations.
I put that into a 4-Ancestors Oracle, along with Yamnaya, Bell Beaker, Corded Ware, Euro Middle Neolithic, Uygur, and a South Asian population with the following:
25 ANE
50 ASE
25 ENF
Which is one of the hypothetical scenarios for the first mixing of ASE with Asians/SW-Asians. In this scenario, a 50/50 ANE/ENF population mixes with 100% ASE. Who knows, it could be another combination/scenario, we won't know without ancient South Asian DNA.
In any case, modern South Asians (from Northwest India) get modeled as 75% of the pre-Yamnaya SC-Asian population and 25% of this ancestral placeholder South Asia. It could be this population or its relatives mixed multiple times with South Asians.
This population is also close to Gedrosian/Baloch component which is why it comes out as Balochistan in Oracles and why some are confusing it for a West Asian population. The ANE is too high, the ENF too low, and WHG too high for West Asia. It's something southeast of Yamnaya towards Central Asia is my guess. If it's real it would've been the product of multiple admixing events between West Asian-like, Steppe-like, and HG-like populations in who knows what order. That's probably what happened with Yamnaya as well, I doubt there was one actual real SC-Asian population with the above profile except perhaps as one of a series of populations emerging from a constant meeting/mixing between Central, West, and South Asia.
FWIW, that would make modern South Central Asians (Pathans, etc) appear as 35-40% Yamnaya if they're 75% of this hypothetical SC-Asian population which would come out as 50% Yamnaya in Admixture, being a parent of it, so that's three 25% chunks of a South Asian admixture profile appearing to be half-Yamnaya = 37.5%, despite having little to no WHG otherwise. At least that's what I assume is meant by people referring to the "non-EHG half of Yamnaya" and Admixture recognizing it and clustering around it.
I generated straight Yamnaya estimates using K8 components for South Asians and besides Haryana Jatts and some Afghan Pashtun who had enough WHG to register 30-40% Yamnaya (and forming a continuum of sorts to Tajiks), everyone else was in the teens or 20s.
For example:
Afghan Pashtun Kandahar: 15.68% South Indian, 32.77% Yamnaya, 49.04% West Asian.
Haryana Jatt: 22.69% South Indian, 39.09% Yamnaya, 33.48% West Asian
Punjabi Jatt Sikh: 26.85% South Indian, 27.26% Yamnaya, 44.36% West Asian
Nepal Brahmin: 31.47% South Indian, 27.06% Yamnaya, 34.20% West Asian
Balochi (an actual individual): 28.82% South Indian, 5.17% Yamnaya, 64.58% West Asian
West Asian emerged "naturally" as leftover ANE/ENF in a 35:65 ratio.
These wouldn't reflect shared genetic drift/ancestry in the way admixture would.
Correction: That was with K7 numbers, not K8. With K8, Yamnaya peaks at 21% in Haryana Jatts.
The WHG estimate of calculators like HarappaWorld are around halfway between K7 and K8.
That's one of the consequences of Admixture that's not so helpful.
@Shaikorth
"Yamnaya weren't 50% EHG in the K9 experiment. Their EHG-like parts were split between WHG, EHG and Central Asian."
I didn't find Yamnaya in the K9 spreadsheet. Where did you see that? My assertion of Yamnaya being 50% EHG came from the other measurements (in Haak et al. 2015, etc...)
EHG had 0% Central Asian, though. So we can assume that Yamnaya in that test would have at least similar levels of EHG and Central Asian. Yes around Europe the Central Asian dominated clearly.
"So there's no real contradiction and the Yamnaya estimates for Europeans in this test do seem to match pretty well the Haak ones from table S9.27."
They do match the ones in Haak because both assume that Yamnaya is the only source of ANE in Europe. Since we don't have other genomes to disprove it, it's ok as an ongoing research. But I think we have many clues that make it clear that some important pieces are missing and the final picture will be quite different.
And to elaborate with some numbers on the Pontic Steppe irony I mentioned above:
In K8, Yamnaya is about:
35/35/25/5 (ANE/WHG/ENF/SE)
Ukrainians about:
17/45/36/2
To reduce the ANE from 35 to 17, it would imply a 50% mix with a population with this profile:
0/55/45/0
But such population just didn't exist anywhere in post-Yamnaya era. Whichever population decreased the ANE by half while increasing the WHG ancestry, had to have some 15% ANE. Which implies almost full replacement of Yamnaya people in Ukraine (by someone similar to Poles).
The lack of Middle Eastern in Basques is very noticeable. Sticks out from all neighbours.
@David
Dhanyavad.
@Alberto
We are going to get Greek aDNA?
The ADMIXTURE tests don't duplicate Haak fits because the clusters aren't based on formal testing. Yamnaya in K8 has almost as much ANE as EHG does, and certainly does not get 50% EHG in K9 even though it is possible as a fit based on formal testing. I'd wager the Central Asian was more dominant in Yamnaya because S-C Asians have significant Yamnaya ancestry and no K9 EHG.
The range for EHG ancestry in Yamnaya depends on references, according to Lazaridis it's 32%-61% in the successful fits they tried. In any case the Haak model can't find out the true level of Yamnaya ancestry in Europe because in their method Yamnaya can be replaced with EHG and increased EEF.
As for Ukraine, the place has indeed gone through many post-Yamnaya population replacements by other steppe peoples and then by Slavic colonists so indeed there was a replacement by someone like Poles, a few centuries ago. But the preceding population probably resembled Crimean Tatars who don't have Yamnaya levels of ANE.
@Nirjhar
There will be this conference about the Aegean Palatial civilizations, I think on March 26th. Chad Rohlfsen will attend, I think. So we might get some info about it fast.
http://eurogenes.blogspot.com/2015/02/a-couple-of-aapa-2015-abstracts-to-blow.html
(We still don't know what kind of genetic analysis they did, but the abstract makes it sound exciting).
@Shaikorth
But Yamnaya samples were not run in K9 (unless I missed that). Of course they would not get 50% EHG, because even EHG themselves only get 75%.
But I doubt they would get 50% Central Asian either. Because the part from the other population would go also to ENF, WHG and SE.
So I guess that Yamnaya would have SIMILAR levels of Central Asian and EHG (difficult to say exactly, but in that ballpark).
Northern Russians do get similar amounts of both, but from there the EHG starts to drop rapidly. Both EHG and Central Asian were completely absent in European Middle Neolithic. So if they both came only from Yamnaya, the levels should stay in the same proportions. But they don't, central Asian stays, EHG drops. It's just an experiment, I know, but it does give a hint that matches many others.
About Ukraine, yes, I know the population movements are more complex, it was just a simplification to say that even there the original Yamnaya population mostly went away (though not their genes, that still amount for some good double figure percentage - but not sure if double figure for EHG ancestry).
David,
Can do some Baalberge samples, plus NE7, CO1, BR1, and Beakers? Thanks!
Well David could run one Yamnaya through K9. I figure they'll be more Central Asian. The North Russian issue can be seen this way: anyone living north of the steppe in the regions that weren't reached by Yamnaya and were at best peripheral to Corded Ware has a greater chance of having ANE from EHG survival, so in Western and Central Europe the portion of ANE coming from the steppe migration should actually be greater.
Yes, but no matter if Central Asian is higher than EHG in Yamanaya, the thing is that EHG drops down to 0% in many parts of Europe (mostly south), while Central Asian stays mostly stable around 10-15%.
The methodology seems sound, appreciated.
Re. Udmurts' peak: they seem to be about the main non-Atlantic pop. with lots of red hair. On the other hand their Y-DNA is almost totally N and they often enough display Siberoid or Mongoloid physical traits.
"Pre-Yamnaya seems to be a mixture of Near Eastern Neolithic and WHG".
If it'd be a mere generic mixture you'd see it probably as such dislocated two components. More likely it is the main founder effect of Thessalian Neolithic and that's why it is homogeneous (however it does seem like a small fraction of it tends to erratically melt with "Middle Eastern"). Anyhow inter-component Fst distances can be used to estimate the relative affinities.
Why didn't you include any sample from Gökhem, Corded Ware or Bell Beaker? I would have liked to see how they'd behave.
Davidski,
Awesome! Can you run this test for the ones that are interested and send a mail with the results?
@Alberto: "But such population just didn't exist anywhere in post-Yamnaya era. Whichever population decreased the ANE by half while increasing the WHG ancestry, had to have some 15% ANE. Which implies almost full replacement of Yamnaya people in Ukraine (by someone similar to Poles)".
Or admixture of early Yamna intruders with a mixed ENF+EHG (=WHG+some ANE) on their way to Poland and the Balcans. For example a base of Dniepr-Don peoples (surely close to EHG) and Cucuteni-Tripolje ones (surely ENF in essence) would do the trick.
Not sure but this would work well, I believe.
Central Asian is also the main, if not only, "ANE-related" component of S-C Asian IE speakers. So the way it could have gone is that South Europeans without EHG in K=9 just don't have EHG beyond what the steppe migrations brought. There could also be overlap.
Maju, the Pre-Yamnaya does very much seem a mix of Near Eastern and WHG. ADMIXTURE can give robust clusters to mixed populations even at low K's (Sardinians and Karitiana are good examples), and you'll need formal testing to figure out how things really are.
@Graham Little: yes but to be fair there seems to be another such pole of extreme low ME component: around the Baltic (Lithuanians, Swedes, some Poles and the occasional Russian).
This may help explain the cross shape that European PCAs tend to adopt: the Russian-Sardinian polarity could relate to the "Yamna" component, while the Basque-Eastern Med one surely relates to the "Middle Eastern" one.
@Saikorth "ADMIXTURE can give robust clusters to mixed populations even at low K's"...
I know. They are still usually discernable via the Fst table that ADMIXTURE provides, at least if Fst distances are marked enough, as happens among inter-continental pops. Not sure if possible here as I haven't seen the corresponding components' Fst table.
"... the Pre-Yamnaya does very much seem a mix of Near Eastern and WHG".
Possibly something of the like but why does the cluster form so naturally? Probably because it is homogeneous across the continent. And this homogeneity suggests a founder effect. Founder effect that I would relate to Thessalian Neolithic (where ME+UHG mixed per Lazaridis).
In fact the figures of {pre-Yamnaya + ME} are not very different from the EEF figures of Lazaridis, while the WHG figures seem to largely correspond to {Yamna - Lazaridis' ANE + other WHG}. At least on preliminary cursory look - am I wrong?
"This may help explain the cross shape that European PCAs tend to adopt: the Russian-Sardinian polarity could relate to the "Yamna" component, while the Basque-Eastern Med one surely relates to the "Middle Eastern" one."
However, Yamnaya are not beyond modern European variation in an European PCA, where the Basque-East Med dimension exists. In typical West Eurasian PCA's they are, but there the dimension that separates them from Europe is not Sardinia-Northeast Euro but Sardinia-West Asia/Volga-Ural and there is no Basque dimension.
Here's two Yamnayas in Euro-only PCA's I asked David to make, where a Basque-related dimension is present (ctrl+f to pinpoint).
https://drive.google.com/file/d/0B9o3EYTdM8lQTnJSeXFucko3ZUU/view?pli=1
https://drive.google.com/file/d/0B9o3EYTdM8lQODRhZDFieEVieDQ/view?pli=1
The pre-Yamnaya cluster peaks in Iceman at near 100%. I have no idea if Oetzi was pure Thessalian neolithic. The component does seem to be a peculiar sort of EEF (WHG+Near Eastern) cluster.
@Maju
"Or admixture of early Yamna intruders with a mixed ENF+EHG (=WHG+some ANE) on their way to Poland and the Balcans."
No, including EHG in the equation wouldn't work (if they mix to such degree with NE pop to lower so much the ANE, they would also lower the WHG). You need to both rise WHG ancestry and lower ANE ancestry. Only a Loschbour-rich population can do this 2 things at the same time.
Re: Pre-Yamnaya, it matches exactly Ötzi (Iceman).
@Maju
Related to the above EHG vs. WHG in the Pontic steppe, it's also an irony that nowadays EHG ancestry is only significant in Uralic and Altaic speakers.
Thanks David for running this:
HRP0393 Haryana Jatt in Yamnaya K6
Yamnaya_related 0.396573
WHG_extra 0.027646
ENA 0.143771
Middle_Eastern 0.410162
Pre-Yamnaya 1E-005
Sub-Saharan 0.021838
The extra WHG is interesting. Have you run a zombie of this Yamnaya through K8?
@Shaikorth
"Central Asian is also the main, if not only, "ANE-related" component of S-C Asian IE speakers. So the way it could have gone is that South Europeans without EHG in K=9 just don't have EHG beyond what the steppe migrations brought. There could also be overlap."
You mean that a later migration brought more Central Asian without EHG to Southern Europe?
I think it did, but why "later"? That would have doubled the Central Asian (and still might not bring to 0% the EHG). It brought it "instead" of Yamnaya. Just makes more sense.
level of homozygosity of the processed diploid genomes -
from the best quality to the worst quality:
I0118.txt 0,738929924
I0100.txt 0,740551005
I0408.txt 0,740769552
I0099.txt 0,741301107
I0406.txt 0,741994462
I0443.txt 0,753786975
I0061.txt 0,758099179
I0112.txt 0,760614205
I0104.txt 0,771278813
I0231.txt 0,771449115
I0103.txt 0,773434461
I0412.txt 0,775065963
I0054.txt 0,775670147
I0172.txt 0,808732446 (this was the biggest BAM, but...)
I0410.txt 0,840994437
I0047.txt 0,882935478
Not a later migration necessarily. I think source for ANE in South and Central Europe could well be the same, just diluted enough in the south that there the EHG component no longer shows in the calculator (and that is assuming Yamnaya in K=9 have any in the first place).
But what's the logic in 2 components starting at a given level, and one of them diluting and the other one remaining at stable levels?
Unless the mixing population already had Central Asian (but that brings us to the same conclusion of different migrations bringing Central Asian).
Yes, it's all assuming Yamnaya had EHG in K9, but if Yamnaya was some 50% EHG it's not a wild assumption. I'm just going for the most likely scenario (with all the caveats of K9 being an experiment not based on real ancient genomes, of course).
Srkz had a IBD map of Mal'ta up. It clearly shows Udmurts as well:
http://i.imgur.com/YquqcDj.png
"But what's the logic in 2 components starting at a given level, and one of them diluting and the other one remaining at stable levels?"
It's something that happens commonly in ADMIXTURE. Same happens with the K15 "Amerindian" in Yamnaya or MA-1. We know European are admixed with them but show no "Amerindian". Shared ancestry and admixture is very easily hidden under components. Two populations sharing no components doesn't equal lack of shared admixture.
"Srkz had a IBD map of Mal'ta up. It clearly shows Udmurts as well:"
Yes, it looks Yamnaya-like. Unfortunately, the quality of the Mal'ta genome is very bad, homosygosity level is 0,94. All long IBD-segments are destroyed.
Well, yes, it can happen. Especially when comparing ancient genomes with modern populations that may have drifted in one specific way. So yes, the evidence is inconclusive.
But it also matches the fact that the south east of Europe could hardly be explained with Yamnaya being the only source of ANE (though again, comparing with modern samples, so also inconclusive).
Let's see if that conference about the Aegean civilizations does shed some light with some real data about this.
@Srlz
Both hotspots on that map are Udmurts, shown here to have a lot of Yamnaya, and Ket, who IIRC had a lot of ANE.
But true. We should keep in the warning mind it could be (partly) artefacts.
@Shaikorth
At K=20, when WHG becomes a separate component in the Yamnaya paper Loschbour almost looks EHG, the HG part of Stuttgart becomes EHG rather than WHG and while all prehistoric populations show both Yamnaya EHG (darkblue) and WHG (grey) all modern European populations only show darkblue. And that is almost certainly not true as the same paper shows.
@Shaikhorth
"The pre-Yamnaya cluster peaks in Iceman at near 100%. I have no idea if Oetzi was pure Thessalian neolithic. The component does seem to be a peculiar sort of EEF (WHG+Near Eastern) cluster."
Was the other 4 x G2a from LBK_EN ( haak paper ) found together in Germany, who have same age as iceman used in this test?
Would they be Thessalian as well?
The ADMIXTURE tests don't duplicate Haak fits because the clusters aren't based on formal testing. Yamnaya in K8 has almost as much ANE as EHG does, and certainly does not get 50% EHG in K9 even though it is possible as a fit based on formal testing. I'd wager the Central Asian was more dominant in Yamnaya because S-C Asians have significant Yamnaya ancestry and no K9 EHG. - Shaikhorth
@David, not to be rude but why are you doing something the researchers have already done? Is the purpose of this post to estimate the yamnaya admixture in non europeans since Haak only did one for europeans?
@Davidski,
This is freaking incredible!!! Great Job. Can you get results for any of us who are in your database in this test?
Did Middle Neolithic genomes trigger the pre-Yamna cluster? Who triggered the Middle Eastern cluster?
@ Alberto
"It's is Yamnaya related, but actually to the non-EHG part of Yamnaya."
So then, is it merely picking up whatever this "West Asian' stuff is ?
@ David
"This is a very peculiar component that is virtually missing from Neolithic Germany, Hungary and even the Copper Age Alps."
David, whilst I have nowhere near as much technical skill as some, I think I can see the wood for the trees. And that it is, I am still a little suspect of the underlying assumptions of the otherwise flawless calculations. So what you're calculating/ running is the problem.
This 'Yamnaya component' is actually not entirely novel- at least compared to the SHG type samples which have, both, "ANE", and "EE/ North Sea/ Baltic" (Whatever component you wish to use, which otherwise gives the same result anyway based on the same data) In fact, even Labschour and Motala have "EE/ Baltic" in addition to Atlantic / North Sea. So really its just a different presentation of the same result of the same data !
What Im saying is many of the BB's just look like Motala-like groups, with much of the "West Med" removed, a touch more EE and a new "west Asian' element.
I don;t understand why the study modelled itself such that ANE could only have come from Yamanaya. It seems like there is no reason for this, other than the constraints of the hypothesis the study seeks to prove- the steppe hypothesis. Not only is this a perplexingly post hoc analysis, but is contrary to the historical-archaeological reality we'd expect from the real evidence on the ground. and No-one has explained why?! Just saying.
* What remains to be seen is what the situation was like in Poland. I know we have intimations from recent Lengyel and TRB mtDNA studies, but they haven't sampled more proximate GAC and "epi-Neolithic" (ie residual Mesolithic) groups from the Baltic Sea littoral. Whatever the case, a westward migration to Germany is undeniable, but to me , i'd be surprised if the results don;t just show this to be a relative "shift" , ie a an internal european migration from anywhere east of the Vistula
@ Alberto
"Which implies almost full replacement of Yamnaya people in Ukraine (by someone similar to Poles)."
Yes, i think we've seen that already based on mtDNA data (?) Combined this with the fact that the Samaran samples left few descendents locally, to me it yet further raises the question of how the steppe set off such population replacements when its people couldn't even linger on in their own environment.
@ Shaikorth
I suspect you're analysis is constrained by the need to fit all the data into an 'out-of-the-steppe model' That model is wrong.
****
I think i know what we're actually seeing with all this data. Its actually rather simple if you know the prehistory of the steppe. And its something @ Alberto has already alluded to......
Does anyone have an opinion what high Yamna-related ancestry in Central-south Asians means? Anyone think it is real Yamna-like ancestry instead of only shared ANE-affinity?
Krefter,
Really hard to say at the moment. My opinion is that it represents steppe migrations to South Asia from east of the Urals, and thus not directly from Yamnaya.
Alberto,
Here are some Bell Beaker f3 stats. I'm now using the inbreed: Yes flag, which is what I should've done before.
Spain_MN Yamnaya Bell_Beaker_LN -0.007705 0.001207 -6.385
Germany_MN Yamnaya Bell_Beaker_LN -0.007757 0.001337 -5.801
HungaryGamba_CA Yamnaya Bell_Beaker_LN -0.007382 0.001838 -4.017
Krefter,
This is an interesting question. For what it's worth, the pattern here is somewhat distinct from ANE levels. Pashtuns have more ANE ancestry than Pamiri Tajiks, but less Yamnaya-related ancestry. Also, ANE has a strong peak among the HGDP South Central Asians, and no population in Europe really approaches that. By contrast, most Northern Europeans have higher Yamnaya-related ancestry in comparison to South Central Asians.
Basically, this is something different from pure ANE ancestry.
A side note, but if the Yamnaya cluster is tracking PIE-related ancestry, it's giving a lower bound for actual Indo-Iranian ancestry among South Central Asians. If we had aDNA from the people who brought Indo-Iranian languages to South Central Asia, I think we will find that people like Pashtuns are more like 60%-80% steppe Indo-Iranian, while Pamiri people are more like 70%-90%. In other words, we might find that there was something close to complete population replacement in the region. But that is just speculation, until we get those genomes (if we ever do), along with aDNA from the Indus Valley civilization+BMAC.
Davidski
"My opinion is that it represents steppe migrations to South Asia from east of the Urals, and thus not directly from Yamnaya."
How do you know what's the chicken and what's the egg ?
@Saikorth: Thanks for the graphs. And effectively Yamna corresponds with the "Russian" polarity (ok, rather Finnic, but roughly same thing), as I expected, vs. the Sardinian one. On the other hand in that particular plot, the Basque polarity is shared by Brits and Scandinavians, something I had not seen before, so I'm wondering what it means. A simple answer could be the Atlantic element I expect to be eventually detected but nothing clear so far. It'd be interesting to see how Gok samples and other ancient ones behave in that Euro-plot, really.
"The pre-Yamnaya cluster peaks in Iceman at near 100%."
And Spain_EN (93%).
Then comes Neolithic_Hungary (also ENE I believe) with 83%, Esperstedt (MNE) with 77% and finally Stuttgart (ENE) with 74%.
Modern Sardinians overlap well with Stuttgart, while Basques have close levels of Neolithic-like "pre-Yamnaya" but rather than showing "West Asian" as secondary component, show "Yamnaya" for some reason.
"I have no idea if Oetzi was pure Thessalian neolithic. The component does seem to be a peculiar sort of EEF (WHG+Near Eastern) cluster".
He was "pure something", as he acts as polarity for that component, which is also very high in some Early Neolithic samples. If anything we seem to see an increase of "West Asian" in Central Europe, suggesting a geographic, not chronological, cline of some sort, albeit a counter-intuitive one admittedly: with growing "West Asianness" towards the North (founder effect probably).
@Alberto: "Only a Loschbour-rich population can do this 2 things at the same time."
Fair enough, I guess.
@Colin: because he feels like. If I had the time and means I'd also be doing my own runs, with different sampling strategies and what-not. It's important to see if results can be replicated independently for example or how much they vary with somewhat different approaches. Just the usual... genetic statistical analysis is not rocket science but it can benefit from getting some of that critical contrast of methods and results that has made space science so paradigmatic of excellence. Anyhow different perspectives increase accuracy in perception, something you can easily when you try to do things with a patch on one eye.
Maju,
"while Basques have close levels of Neolithic-like "pre-Yamnaya" but rather than showing "West Asian" as secondary component, show "Yamnaya" for some reason. "
It's quite clear surrounding Euro pops gave Basque eastern ancestry. No one in west Asia fits the bill.
@Seinundzeit,
"Basically, this is something different from pure ANE ancestry."
There's nothing representing south Asian though. I suspect there's significant Yamna-type ancestry in south-central Asia. I don't think anyone has found an accurate way to distinguish between Yamna-ANE-ENF-WHG and native-ANE-ENF though.
"A side note, but if the Yamnaya cluster is tracking PIE-related ancestry, it's giving a lower bound for actual Indo-Iranian ancestry among South Central Asians."
Of course that's the case. A Sumerian genome will prove that at least Kurds and Iranians have significant ANE-heacy non-near eastern ancestry. It's hard to imagine an ancient ethnic identity and language has little to do with genetics anywhere in the world.
Sure, you can give examples like Spanish is from Latin, but there was nothing like the Roman empire or modern countries in most of the ancient world.
''"A side note, but if the Yamnaya cluster is tracking PIE-related ancestry, it's giving a lower bound for actual Indo-Iranian ancestry among South Central Asians."
Nope.
''Of course that's the case. A Sumerian genome will prove that at least Kurds and Iranians have significant ANE-heacy non-near eastern ancestry. It's hard to imagine an ancient ethnic identity and language has little to do with genetics anywhere in the world.''
oh there will be many more to topple your world trust me Krefter...
'' It's hard to imagine an ancient ethnic identity and language has little to do with genetics anywhere in the world.
Sure, you can give examples like Spanish is from Latin, but there was nothing like the Roman empire or modern countries in most of the ancient world.''
Yes quite can be but also not can be! and obviously its too early to Draw any conclusion specially for Asia IMHO.
@ Nirj
The yamnaya figures are inflated and blatantly incorrect
They don't know what they're actually seeing
@Krefter: Neither Spaniards nor French are that strongly "Yamnaya" (in this run not certainly) to cause such an effect. Basques are in the order of 20-25% "Yamnaya" (i.e. Finnic-like per Chad's plot), Spaniards and South French are in the same range, while (North?) French are just in the 25-35% range. Also all them have much more "Middle East" than (most) Basques, what seems a detector of the kind of the kind of admixture you propose, marking negative.
There are hidden confounding factors almost without doubt.
@Mike
''The yamnaya figures are inflated and blatantly incorrect
They don't know what they're actually seeing''
Then why create such a component and ADMIXTURE? if a better attempt can be made?
I'll rephrase ; as I realize that Im being self contradictory .
The figures are correct ; but they're not actually detecting "yamnaya " intrusion rates ; which were actually minimal
Mike, OK:P, So ''EHG received much but Diffused less''?
Nirjhar
Yes.
I think people have made some good points here which reuire highlighting for the sake of the 'big picture'
* Davidski said "So it's not implied here that anyone who has it also has Yamnaya ancestry"
* Gill said "perhaps as one of a series of populations emerging from a constant meeting/mixing between Central, West, and South Asia."
What we're seeing is complex series of admixtures which culminate in the late neolithic. Yamnaya was but one region/ "culture" involved in this process, and not the ultimate progenitor. These 'events' were not unilinear, and spanned from eastern Europe to central Asia. We know they took place in the north Ponto-Caspian region (ie the steppe). But did they also take place to the south ? Ie the south caspian corridor from central Asia to anatolia to Balkans ???
Prior to all this, as we know, the seppe was still very much EHG type genetically; demographically sparse, and culturally peripheral.
So the real events are far more complex. That is clear. Yet, people still try to hammer it into a Yamnaya-centric narrative, for various reasons ........
@maju
In Northern France, the middle Eastern is propably linked to the LBKs.
We exactly see the same thing with the caucasian in other calculators.
We have a lack of data, this component certainly increases from west to East.
From Brittany to Alsace.
I had a chance to look and compare K15. One thing must be taken into account. K6 is modeled around Yamnaya I0429. The oldest dated R1b sample is within roughly 1 mile from this sample 53+/-_Lattitude 50+/-_Longitude https://docs.google.com/spreadsheets/d/1QPTmyarOBBEZfXnLI5L64ueJNG34jgy4QgQ_1nSYtnM/edit#gid=917906623.
modled for K6 99% related Yamnaya We know the time frame is roughly 7650+/- & 5354+/- between these 2 R1b samples with the latter being R1b Z2103+L584- . The oldest R1b sample has the highest reading of any known Eurogenes K15 Eastern European component; plus some North Sea [ represented by Norwegian]plus some Amerindian[Katriana component] Baltic[Lithuanian]The younger R1b sample is from a mile away and differs by a spike in West Asian[represented by Georgian or Proto-Kartvelian] and South Asian[Uttar Pradesh?]
Yes, the 100% Yamnaya genome looks like an extreme example of a Yamnaya sample. You can see that on the K8 PCA.
http://eurogenes.blogspot.com.au/2015/03/bell-beaker-corded-ware-ehg-and-yamnaya.html
Any chance of releasing this calculator on GEDmatch? I hope you can include some more South Asian ethnic groups.
Mike Thomas,
My analysis is primarily based on the Haak results, and I do think non-steppe related EHG survival is probable in North Europe, especially Northeast Europe. So no, not everything must have come from the steppe.
"What Im saying is many of the BB's just look like Motala-like groups, with much of the "West Med" removed, a touch more EE and a new "west Asian' element. "
Are you sure you didn't mean some other group than Motala? BB's actually have much lower East Euro than Motalas, but because of increased Near Eastern ancestry they have higher Atlantic, and also West Asian and West Med which the Motalas lack.
"I don;t understand why the study modelled itself such that ANE could only have come from Yamanaya. It seems like there is no reason for this, other than the constraints of the hypothesis the study seeks to prove- the steppe hypothesis."
They modeled Europeans with zero Yamnaya and all ANE from EHG survival too, as can be seen in tables S9.25-S9.27. However, "no steppe contribution" is even more unlikely for ANE than "all steppe contribution".
@Helgenes: All I said is that the "ME" component works as evidence that excludes normal contact admixture with neighbors being at the origin of the "Yamna" component in Basques. Because neighbors (many types of Spaniards, Gascons and French) have "ME" and most Basques do not, while "Yamna" is found at similar frequencies in all these groups.
Not sure how your comment can be related to this.
Shaikorth
Thanks for your explanation
I agree with what you said. But I wonder: how secure should we be with the way we extrapolate admixture rates backward from the proportionality of changes we see in our timeline of genomes ?
""However, "no steppe contribution" is even more unlikely for ANE than "all steppe contribution""
Well of course . I've never suggested what would be an absurd proposition.
I'm rather suggesting that pre-Yamnaya ANE has been nevertheless Underestimated simply because it was deemed "unlikely" by the authors to have been the primary source of ANE in Europe (with which I agree); yet am surprised that they didn't model for SHG type sources to have also contributed, on the one hand. On the other hand; I think the point being missed is that yamnaya was more the vector than the primary catalyst for the demographic changes seen, in addition to lack of resolution as to how much, if any, this yamnaya -like intrusion actually and specifically derives from yamnaya itself.
@ maju
Yes indeed, my comment is a little off topic, I am sorry.
I try to find the best marker to differentiate the LBK and the cardial.
The reason, the neolithic in Normandy is Atlantic and Danubian
The local archaeologists still tend to focus on the second.
@Mike
"I'm rather suggesting that pre-Yamnaya ANE has been nevertheless Underestimated simply because it was deemed "unlikely" by the authors to have been the primary source of ANE in Europe (with which I agree); yet am surprised that they didn't model for SHG type sources to have also contributed, on the one hand."
Yes, Motala ANE has basically been taken out of the picture for being unlikely source of anything. However, there's something that no one commented about in the paper:
Figure S7.7 (page 66): While CW shares a bit more drift with EHG then with Motala, the opposite happens with Bell Beaker, Unetice and Halberstadt. The case of Unetice is specially striking, since it really shares significantly more with Motala than with any other population.
I'm not really sure how meaningful it is, but at least it raises some question as whether Motala-like populations could have been around Poland, Czech Republic, Belarus,...
@David
Thanks for those f3 stats, something was puzzling us with those Bell Beakers :) Related to what I said above, could you try these?
Spain_MN Motala_HG Unetice_EBA
Spain_MN Yamnaya Unetice_EBA
Thanks.
Here you go...
Source 1 Source 2 Target f_3 std. err Z SNPs
Spain_MN Motala_HG Unetice_EBA -0.003758 0.001045 -3.598 154775
Spain_MN Yamnaya Unetice_EBA -0.009940 0.000965 -10.296 156311
Thanks!
Those stats with the Dinka outgroup seem a bit misleading. I found the Fst distances table more informative.
Unetice - EHG: 0.034
Unetice - Motala_HG: 0.051
Unetice - Yamnaya: 0.012
Also it confirms that BB and Unetice are much closer to modern Europeans than the older Neolithic samples.
Spanish - Spain_MN: 0.020
Spanish - Bell Beaker: 0.006
Spanish - Unetice: 0.007
@Mike
Yes, there might be something there with Motala like populations. However, the information is contradictory (see David's f3 stats and my following comment). It's difficult to say which stats are more correct. Maybe a mix of both (as you always point out, these processes are complex, so statistics can often be misleading when trying to draw a simple picture).
That's my point Alberto. It doesn't have to be one or the other ; but could be both. We shouldn't limit our exploration based on small differences in statistical probability
Mike, Is that you in the Pic?
Yeah bud.
Some preliminary results using qpAdm...
Corded Ware
WHG 0.117
LBK 0.190
Yamnaya 0.694
Esperstedt_MN 0.270
Yamnaya 0.730
Yamnaya
EHG 0.514
Armenian 0.486
EHG 0.559
Iraqi_Jew 0.441
Icelanders
Motala_HG 0.000
Germany_MN 0.526
Yamnaya 0.474
That looks promising.
Can you try Yamnaya as EHG + BedouinB + Kalash? Haak method can fit it as just EHG+BedouinB, and if it is a simple EHG/Near Eastern mix it shouldn't get additional Kalash.
This is a pretty good fit.
EHG 0.523
BedouinB 0.222
Kalash 0.255
Does it give a range of possible fits or just the best one with given references?
Next best is this, but it's way behind the first option that I just posted.
EHG 0.653
BedouinB 0.347
Kalash 0.000
The other five options are terrible fits.
Alright. Then it looks pretty decent at gauging things like Motala survival.
If
Motala_HG 0.000
Germany_MN 0.526
Yamnaya 0.474
is a better fit than one that gives Icelanders Motala as well, we could consider that evidence against ANE survival caused by SHG types in NW Europe.
That's cool. Is it limited to a number of populations? If 4 are possible, and if it's easy to try, would it be possible to run this one?
Tuscan:
LBK_EN
Loschbour
Yamnaya
Lezgin
Good idea, although Lezgins have considerable overlap with Yamnaya because of their particular mixture.
What do Tuscans as these look like:
LBK_EN
Loschbour
Yamnaya
BedouinB
LBK_EN
Loschbour
Yamnaya
Dai
Top 4. As far as I can see from the output, none of these are great fits, but they work.
LBK 0.488
WHG 0.000
Yamnaya 0.000
Lezgin 0.512
LBK 0.742
WHG 0.137
Yamnaya 0.121
Lezgin 0.000
LBK 0.803
WHG 0.000
Yamnaya 0.197
Lezgin 0.000
LBK 0.692
WHG 0.308
Yamnaya 0.000
Lezgin 0.000
"Good idea, although Lezgins have considerable overlap with Yamnaya because of their particular mixture."
Yes, I know. The point I'm trying to figure out is if Southern Europeans (basically ancient Italians and Greeks) derived from Yamnaya or from the "other population" that was part of Yamnaya.
For the results, Tuscans don't seem to like Yamnaya much, but Lezgins are not the best fit either (as source of ANE).
I think that ancient DNA is the only way to know, because modern pops are more admixed and drifted, so it's all too speculative.
Thanks David.
Shaikorth, the first one was a disaster, but the second worked I think. Keep in mind I've just started with this thing.
LBK 0.742
WHG 0.137
Yamnaya 0.121
Dai 0.000
LBK 0.717
WHG 0.275
Yamnaya 0.000
Dai 0.008
Looking at some old f3 stats for modern populations, the best fit for Tuscans as mix of Sardinian + other was with Gujarati3:
Tuscan;Sardinian,Gujarati3 -0.00154699 9.27455e-05 -16.6799
So using Gujarati3 instead of Lezgins could work. But I'm still sceptical as to how much can we trust results when mixing ancient and modern samples.
Ah... Interesting exercise to recreate the methodology of Haak's residual modelling n=4 (plus African ancestry) as close as ADMIXTURE can.
Here's a quick triangle plot based on this K6 which I thought was interesting to look at - http://i.imgur.com/xTIxZQj.png. These clusters are probably less equidistant than the the K8 ones, so maybe the genetic distances from PCA map a little less to this PCA, still fairly interesting to look at.
Btw, David could you rerun these f3 stats with the Inbreed parameter set to yes -
http://textuploader.com/mu6r? And the D-stats http://txt.do/muly. Thanks.
This qpAdm seems interesting (essentially it's the Haak Modelling, automated, right?). Right now can't really think of any great tests for it though that Shaikorth hasn't pulled up - maybe Armenian as LBK_EN Yamnaya and GujuratiC/Sindhi or BedouinB EHG and GujuratiC/Sindhi? What about EHG as WHG MA1?
Matt, the D-stats are the same, because they're not affected by inbreeding. Here's the other stuff, starting with the more or less successful models.
Tuscans
LBK 0.741
WHG 0.000
Yamnaya 0.140
GujaratiA 0.119
LBK 0.742
WHG 0.137
Yamnaya 0.121
GujaratiB 0.000
LBK 0.742
WHG 0.137
Yamnaya 0.121
GujaratiC 0.000
LBK 0.742
WHG 0.137
Yamnaya 0.121
GujaratiD 0.000
Armenians
LBK 0.860
Yamnaya 0.044
GujaratiC 0.096
LBK 0.821
Yamnaya 0.026
Sindhi 0.153
BedouinB 0.536
EHG 0.096
GujaratiC 0.367
BedouinB 0.449
EHG 0.057
Sindhi 0.494
Loschbour Alberstedt_LN Bell_Beaker_LN -0.005968 0.002122 -2.813
HungaryGamba_BR1 Corded_Ware_LN Bell_Beaker_LN 0.000497 0.002195 0.226
HungaryGamba_BR2 Corded_Ware_LN Bell_Beaker_LN -0.000208 0.001759 -0.118
Spain_MN Yamnaya English -0.002554 0.000679 -3.76
Germany_MN Yamnaya English -0.003466 0.00082 -4.224
Spain_MN Yamnaya French -0.00351 0.000564 -6.224
Germany_MN Yamnaya French -0.004167 0.000723 -5.761
Spain_MN Yamnaya Norwegian -0.001045 0.000678 -1.541
Germany_MN Yamnaya Norwegian -0.001803 0.000814 -2.214
Spain_MN Yamnaya Bell_Beaker_LN -0.007705 0.001207 -6.385
Germany_MN Yamnaya Bell_Beaker_LN -0.007757 0.001337 -5.801
HungaryGamba_CA Yamnaya Bell_Beaker_LN -0.007382 0.001838 -4.017
Spain_EN Loschbour Germany_MN -0.010137 0.002629 -3.856
LBK_EN Loschbour Germany_MN -0.015754 0.002367 -6.656
LBK_EN LaBrana1 Germany_MN -0.012221 0.002532 -4.826
LBK_EN Loschbour Germany_MN -0.015754 0.002367 -6.656
LBK_EN HungaryGamba_HG Germany_MN -0.009827 0.002865 -3.43
LBK_EN HungaryGamba_CA Germany_MN -0.002449 0.00289 -0.847
Balochi EHG Yamnaya -0.004522 0.000997 -4.537
Sindhi EHG Yamnaya -0.002509 0.001004 -2.499
GujaratiA EHG Yamnaya -0.002244 0.001095 -2.05
GujaratiB EHG Yamnaya -0.000647 0.001128 -0.574
GujaratiC EHG Yamnaya -0.000562 0.001133 -0.496
GujaratiD EHG Yamnaya -0.001197 0.00115 -1.041
Yamnaya BedouinB Pathan 0.002364 0.000438 5.4
P.S. Three of the combinations had no data. I need to look into that.
And the EHG modeling didn't work out. This might have something to do with the fact that some of the outgroups have very high ANE levels...?
Regarding new ancient DNA studies. I've found this news on russian websistes: http://infobaikal.ru/news/s179/n127173/
The text contains English translation. Basically scientists from Denmark will try to get ancient DNA from 8000 years old remains.
Another website (http://www.kommersant.ru/doc/2682568)has information only in Russian, but gives photo with facial reconstruction :http://im1.kommersant.ru/Issues.photo/OGONIOK/2015/009/KMO_147099_00024_1_t218_154439.jpg
I understand that this is not European remains, but I think it will be interesting for finding out who were ANE's.
@David
How are Greeks doing? Similar to Armenians?
David,
Your Eurogenes K15 calculator does not seem to be adequate for Yamnaya admixture analysis since the analysis is based on 140,000+ markers.
I just ran Yamnaya through the calculator, and the coverage was only ~60,000 SNPs.
Since the K15's accuracy is dependent on around 140,000 SNPs, Yamnaya's analysis falls short using this calculator.
Are you planning releasing a calculator that uses less markers such as MDLP, to address low coverage ancient genomes?
While Northern Europeans have been swapping components with one another for a long time, Italy has had an extremely strong post-Bell Beaker genetic influx in the form of haplogroups J2 that Northern Europe has not. So, running ADMIXTURE on modern Tuscans, while interesting, does not give a complete genetic picture of Central Italians after the first large R1b influx and before the first large J2 influx. While there is also one single E-V13 from Neolithic Spain, it is not clear when E-V13 really expanded to the high levels seen in the Balkans and Italy. Again, this influx seems to have has little impact on the rest of Europe.
When was the J2 influx into Southeastern Europe? I always thought it was with the first early neolithic farmers and sailors from the Mideast (related to Phoenicians or whoever was in the Levant).
At least that was what the old descriptions of J2b2-M241 used to say.
It appears it came to India around the same time, probably also with a wave of early neolithic farmers. YFull estimates divergence between the branches at 8-9kybp (and having formed around 13-14kybp).
Xinjiang stuff, for someone with access to relay here.
http://www.nature.com/ejhg/journal/v23/n4/full/ejhg2014134a.html?WT.ec_id=EJHG-201504
@Gill
''It appears it came to India around the same time, probably also with a wave of early neolithic farmers. YFull estimates divergence between the branches at 8-9kybp''
Yep.
@ David, thanks, I did wonder that about the D-stats but wasn't sure. Re: f3s, looks like the Yamnaya f3 stat (from the paper) as admixture between populations like EHG and South-Central Asians are negative although not as strong as Armenian and EHG.
Not totally surprised Armenian fits a bit more with Sindhi admixture in rather than Yamnaya, even though Sindhi has some ENA. Armenian as Balochi, LBK, Yamnaya might be even more Balochi...
Remaining one might be Balochi / Sindhi as BedouinB Yamnaya Dai and Balochi / Sindhi as LBK Yamnaya Dai. Dai, in the absence of Onge - although the Kharia people in Haak's dataset might be a better representative of ancient populations of India (the ADMIXTURE gives then the Southeast Asian and South Indian components solely).
@Gill "When was the J2 influx into Southeastern Europe? I always thought it was with the first early neolithic farmers and sailors from the Mideast (related to Phoenicians or whoever was in the Levant).
At least that was what the old descriptions of J2b2-M241 used to say.
It appears it came to India around the same time, probably also with a wave of early neolithic farmers. YFull estimates divergence between the branches at 8-9kybp (and having formed around 13-14kybp)."
It doesn't really matter when it made is way into India... it is missing in all Early and Middle Neolithic samples from Europe, so it is hard to believe that it was hiding in some extreme SE corner of Europe without even a single sample getting swept into places like Spain or Central Europe by other farmers.
@David, another thought -
On a previous post you mentioned
"I'd say an even better solution would be to model the Yamnaya as a three-way mixture between Eastern European foragers, early Neolithic farmers straight from the Near East, and perhaps some sort of Central Asian population very similar to the main ANE-proxy MA-1 or Mal'ta boy"
Is that something we can now try a test of with qpAdm, Yamnaya and EHG, MA1 and BedouinB? Might not work for the same reasons you mentioned the EHG modeling not working out.
EHG+Kalash+BedouinB is a better qpAdm fit than just EHG+BedouinB and that supports David's idea I guess, but wouldn't be surprised if MA-1 produces a weird result.
@Gill
"When was the J2 influx into Southeastern Europe?"
If farmers had two sources, a more maritime Levant one and one from (potentially) much further east then I guess J2 could have been carried on either one.
@Krefter
"Does anyone have an opinion what high Yamna-related ancestry in Central-south Asians means? Anyone think it is real Yamna-like ancestry instead of only shared ANE-affinity?"
I think there was a mega foot herder expansion from somewhere adjacent to the Himalayas and people of partly that same ancestry went to India in mounted form at a later time.
Does the qpAdm give any score to compare between different runs?
For example, for Yamnaya a good fit was:
EHG 0.523
BedouinB 0.222
Kalash 0.255
But if you made another run with:
EHG
Tabassaran
Loschbour
Is there a way to compare which one is a better fit? Or it only gives a hierarchy for a given run?
@ Shaikorth EHG+Kalash+BedouinB is a better qpAdm fit than just EHG+BedouinB and that supports David's idea I guess, but wouldn't be surprised if MA-1 produces a weird result.
Good point; though if Yamnaya fits as, EHG 0.523, BedouinB 0.222, Kalash 0.255 and EHG 0.514 Armenian 0.486, then that seems to kind of imply that the BedouinB 0.222 and Kalash 0.255 (or 0.475 BedKalash) are functioning together similar to the Armenian 0.486 to alter Yamnaya's relationships to outgroups relative to EHG.
Some light comments:
1) I'm really glad to see y'all playing heavily with our data. The data is rich,
and there are very likely phenomena present that our group has overlooked.
2) Be careful not to overemphasize what can be learned from this data from
the MT and Y, especially with deductions about the origin of a mutation.
Laurent Excoffier has shown that it's easy for a mutation to arise in a region where it
is not even present today. The Spanish R1b is interesting but very hard to interpret
by itself. Strange things happen, for instance:
http://www.nature.com/ejhg/journal/v15/n3/full/5201771a.html#close
Note for instance that Yamnaya were probably patrilocal, the horizon is huge,
and our Yamnaya samples are from a small region. One should not try and deduce too much.
from our tiny number of samples from Samara.
3) I'm really glad that Davidski is playing with qpAdm. This is a new program
that I myself am learning how to use. It does make weak(?) phylogenetic
assumptions though. Users have to supply a list of
"left" and "right" populations, and if there were funky migrations between left and right
after the admixture event of interest it won't give the right answer.
4) There's been some criticism on the blog about the populations we studied. Why did we study A not B?
This work is not like ordering a Latte from Starbucks! We can only study samples that are
available to us, and many of these fail to yield useful DNA after screening.
5) There's a new paper by our group and collaborators on selection and phenotype in aDNA.
bioRxiv doi: http://dx.doi.org/10.1101/016477
Nick Patterson
So, Mathieson et al shows Genetiker was correct and the Motala SHG's indeed carried the East Asian EDAR variant. Now the question is whether there was ENA geneflow to Europe at some point before the arrival of Near Eastern farmers (as the possibility was raised in Haak et al) or whether ANE populations carried SNP's associated with East Eurasian morphology these days.
@ Nick Patterson
Thanks for your links and comment!
As far as I can understand, from you point 4, we can expect new results from your labs on ancient human DNA?
@rozenfag
Well we have an aDNA wetlab, and
lots of collaborators, so I sure
hope our recent paper is not
the last word!
... " the neolithic in Normandy is Atlantic and Danubian".
Are you sure there are no Cardial influences there? Also High Normandy was then more related to Brittany and other "Armorica" (hence to one of the earliest centers of Dolmenic Megalithism), while Low Normandy seems rather related to NW France and Belgium (which eventually ends up Danubian but was not early on).
In any case Normandy incidentally hosts the eponym of the La Hoguette culture, more commonly found further East towards South and West Germany, in unclear overlap with LBK, and often argued to be of either Cardial or aboriginal development (or a mix of both - but certainly not Danubian). Later on Danubian advances on NW France but always North of the Seine, not much further south, being unclear how much of the pre-Danubian Neolithic cultures of La Hoguette and Limburgian were assimilated or just wiped off in mass genocide).
I'm open to criticisms of these interpretations, of course.
Shaikorth,
Here's a quote from that new paper...
"The statistic f 4 (Yoruba, Scandinavian hunter-gatherers, Han, Onge Andaman Islanders) is significantly negative (Z=-3.9) implying gene flow between the ancestors of Scandinavian hunter-gatherers and Han so this shared haplotype is likely the result of ancient gene flow between groups ancestral to these two populations."
http://dx.doi.org/10.1101/016477
Yeah, that probably wasn't picked up earlier because Onge and Karitiana were the primary ENA references. I mentioned some time ago I'd like to see things like Han and She ENA references too, just because of a hunch. Apparently not a pointless one.
@ Nick Patterson
Have you looked at the Bolshoy Oleni Ortov samples from Kola Peninsula 1500 BC?
Re: Yamnaya as a modelled combination of populations again - if Pathan 45% ENF and Kalash are more or less like Pathan plus drift, and BedouinB 100% ENF, then Kalash 0.255 and BedouinB 0.222 goes to 11% plus 22%, 33% ENF. Or if BedouinB is 90% ENF, 10% African, then 32% ENF.
Compare to Armenian 77% ENF * 0.486 = 37% ENF or Iraqi Jewish 82% ENF * 0.441 = 36% ENF.
Those all seem to fit with around about as much "Basal Eurasian" effect as I would think it should have based on the Yamnaya D stats David calculated up for me as well.
So maybe we'd see around EHG 0.5 BedouinB 0.35 MA1 0.15? If it works at all.
@Shaikorth So, Mathieson et al shows Genetiker was correct and the Motala SHG's indeed carried the East Asian EDAR variant.
Yeah, despite his general kookiness Genetiker seems to have actually got this one right, and on the pigmentation genetics of Scandinavian HGs.
The derived allele of SLC24A5 seems not actually an agricultural thing, and theories based on that idea are probably wrong.
It's hard to think of a selective pressure that would tie together SHGs and Neolithic farmers while leaving WHGs as an outgroup though. KO1 has one copy of the allele though, and is homozygous, so it's not like WHGs didn't have the allele, just the other ones we have so far (Loschbour and La Brana, who were 1000 years earlier than KO1) did not really have it.
SLC24A2, the "European" only depigmentation variant does appear first in European Neolithic populations, which does make sense.
EDAR is especially interesting because you'd think that if it was in SHG, it should be in EHG, as SHG more or less seem to model as EHG plus WHG.
So if it was in EHG, why not Yamnaya, and if Yamnaya, why not modern Europe and West and South Asia?
East Asian derived EDAR seems like a more or less global gland updevelopment variant - why would that be useful from SHG to East Asia to America, and then *not* in West Eurasians?
The paper: Our data strengthens previous reports of the late appearance of lactase persistence in Europe with the earliest appearance of the allele in a central European Bell Beaker sample (individual I0112) who lived approximately 4,300 years ago. We detect no evidence of lactase persistence in Early Neolithic farming populations like the Linearbandkeramik (LBK), or in the steppe pastoralist Yamnaya, despite their use of domesticated cattle (Figure 2).
So Yamnaya are not the "cow boys". I wonder if that will cut any ice with the people who insisted on a lactase fueled pastoralist expansion because the allele is found from India to Spain? Instead the allele is Bell Beaker first, supporting a link to cattle culture from the West (or wherever Beaker actually came from).
It's interesting how a little adna can overturn a few wrong selective ideas.
The paper: Both the Iberian Early Neolithic and Middle Neolithic samples show evidence of selection for decreased height relative to present-day European Americans (Figure 3A; p=0.002 and p < 0.0001,respectively). Comparing populations that existed at the same time (Figure 3B), there is a significant signal of selection between central European and Iberian populations in each of the Early Neolithic, Middle Neolithic and present-day periods (p=0.011, 0.012 and 0.004, respectively). Therefore, the selective gradient in height in Europe has existed for the past 8,000 years. This gradient was established in the Early Neolithic, increased into the Middle Neolithic and decreased at some point thereafter. Since we detect no significant evidence of selection or change in genetic height among Northern European populations, our results further suggest that selection operated mainly on Southern rather than Northern European populations
So height selection differentiates between Middle Neolithic Germans and Spanish, not just Yamnaya and MN Europeans.
Curious though because Spanish are not that short today, having a similar height (IRC) to English, French, Russians and Swiss.
While the measures of genetic height here show a signal of selective difference between IBS (Iberians 1000 genomes) and CEU (Central European - British Utahans) similar to the one between Iberian_EN and CEU.
I wonder if there might be a link between lactase and short stature in Iberians, where milk consumption tends to give a body weird cow hormones, so to right size the body it tends to pay off to shrink a little? Probably not though.
"Curious though because Spanish are not that short today, having a similar height (IRC) to English, French, Russians and Swiss. "
CEU might resemble the continental North Sea rim more than English in height selection. I haven't seen a height study on white Utahns so hard to say.
Looking at the probably most reliable studies (criteria: measured the subjects' height, no estimations or self-reporting) taken over the last five years, there's unfortunately no Spanish data, but some things may be gathered from the following male height stat:
Serbia: 182 cm (supports a Dinaric Alp selection?)
Montenegro: 183.2 cm (the same?)
Slovenia, Denmark, Finland: 180.3-180.7 cm
England: 175.3-177.8 cm
The data on French and Iberians is older but usually doesn't suggest they are taller than English.
Here's another link to the new paper...
http://biorxiv.org/content/early/2015/03/13/016477
Alberto,
I'm not sure yet how to compare results from different qpAdm runs. I'll look into that.
Matt,
The EHG/BedouinB/MA1 model for the Yamnaya doesn't produce any good fits.
@Matt:SLC24A2, the "European" only depigmentation variant does appear first in European Neolithic populations, which does make sense.
I think you mean SCL45A2 (rs16891982), which both EHG and SHG have in mainly the "white" version (3/4 and 10/14 alleles respectively). I agree, it seems that the first "white" people in Europe were HG's from the North/North-East, not farmers from the South/South-East.
EDAR is especially interesting because you'd think that if it was in SHG, it should be in EHG, as SHG more or less seem to model as EHG plus WHG.
Dave's f4's on Yamnaya show an increased affinity to Dai in EHG, SHG and MA-1, and a decreased one in Stuttgart and K14 - there definitely seems to be some additional ENA in ANE, which may be the source of the EDAR.
@Krefter:
The link you provided to the paper on phenotype in aDNA leads to nowhere.
Google it without the "http://" at the front (or click here).
@ Tobus: I think you mean SCL45A2 (rs16891982), which both EHG and SHG have in mainly the "white" version (3/4 and 10/14 alleles respectively).
Yeah, that's the one - it appears to be like that in Fig 2 (although no EHGs in this paper). Curious as to why they didn't mention SLC45A2 specifically in SHGs in the introduction, rather than "SLC45A2 first appears in our data at low frequency in the Early Neolithic", followed up by "While the western hunter-gatherers of central and southern Europe largely have the ancestral allele at the two major European skin pigmentation loci, the closely related Scandinavian hunter-gatherers have both the derived alleles contributing to light skin pigmentation at high frequency (Figure 2B). Thus, the derived allele of SLC24A5 was common in both the Scandinavian hunter-gatherers and Early European farmers, but not in the geographically intermediate western hunter-gatherers.".
@Shaikorth: CEU might resemble the continental North Sea rim more than English in height selection. I haven't seen a height study on white Utahns so hard to say.
I haven't seen any either. A check they could add here would be to look at Finns in 1000G vs CEU 1000G vs GBR 1000G. I haven't heard there's any statistically significant differentiation between any of these on these height loci, perhaps that would be something they could check out in the supplement.
One more comment re height stuff.
Annoying that many of the differences in the Extended data Figure 5 weren't labeled - a supplement listing them would be useful here. To be fair, they weren't statistically significant differences, but still.
Still check out -
http://i.imgur.com/QGDX7a9.png (I've rescaled and synced up A and B).
For Height Z (score for height differences) Yamnaya-Central MN is around +3, and Yamnaya-CEU is around +3. Suggests CEU and Central MN around the same height.
Difference between CEU and Central MN seems to be as here, around +0.25 Z to CEU (no significant diff.)
Then Corded Ware as 73% Yamnaya, 27% Central MN, then 0.73*3+0.27-0.25, Corded Ware should be around Z +2.1 to CEU.
However, although Corded Ware isn't labelled, the only ancient population other than Yamnaya to have a positive Z when compared to CEU looks around +0.25, non-significant difference. And that population with taller height than CEU could well even be Bell Beaker or Unetice, who I think are also included.
Perhaps this is stretching too far though. Still it would be curious if Yamnaya were selected for taller height than CEU, Central Middle Neolithic for around the same height as CEU, then populations that appear admixed between these two then not appear selected to be taller than CEU, or even closer to Yamnaya, and have no significant difference.
Also wondering if the Motala samples allowed for height score comparisons or not?
@Davidski The EHG/BedouinB/MA1 model for the Yamnaya doesn't produce any good fits.
Thanks, shame we can't tell whether that is real (ENF, EHG plus ANE doesn't work) or somehow due to any issues with MA1 itself as representative for ANE.
"CEU might resemble the continental North Sea rim more than English in height selection. I haven't seen a height study on white Utahns so hard to say."
If there was a significant mixing in England during the industrial revolution of a taller and a shorter population then the ancestors of the CEU may have emigrated beforehand?
CEU aren't pure English by any means, the population history of Utah involves later German and Scandinavian immigration, although the biggest part should be New England colonial.
Early Mormons were apparently quite successful converters in 19th century Denmark and Utah gained a large number of early immigrants that way.
http://www.uen.org/utah_history_encyclopedia/d/DANISH_IMMIGRATION.html
@Matt: Curious as to why they didn't mention SLC45A2 specifically in SHGs in the introduction, rather than "SLC45A2 first appears in our data at low frequency in the Early Neolithic"
Especially since they list Motala at 7.7 kybp and the earliest EN (Starcevo) at 7.6 - technically SHG was first, and at a high, not low, frequency... it does seem a bit of an oversight.
Having rechecked the data I agree with them re SLC24A5 - it was already in high frequency in the ealiest farmer samples (as well SHG/EHG) so a Southern farmer source for this allele is most likely.
I wonder if they're going to publish the calls - I'm going off Genetiker's results and it'd be nice to know if they are duplicated.
@Matt
some guesses
"It's hard to think of a selective pressure that would tie together SHGs and Neolithic farmers while leaving WHGs as an outgroup though."
If two reasons for depigmentation were
1) latitude (for whatever reason)
2) farmer diet
then the first would more likely have developed among archaics implying they'd be the most likely source imo.
If those archaics survived latest in mountainous regions then two widely separated populations might have picked them up separately: say SHGs from Scandi or Urals and farmer ancestors from Urals or Himalayas etc.
.
"East Asian derived EDAR seems like a more or less global gland updevelopment variant - why would that be useful from SHG to East Asia to America, and then *not* in West Eurasians?"
One possibility would be if it duplicates a west Eurasian version of the same thing e.g. say two genes developed in separate regions that provide an advantage via breast milk but only can operate at a time.
.
"So Yamnaya are not the "cow boys"."
Atlantic cowboys.
The difference may somehow be related to which herd animals were dominant in particular bioregions: mares, goats/sheep or cows.
If mares were the most cost/effective in the east for some reason, goats and sheep in the south for some reason and cattle only in the northwest (due to Atlantic rainfall).
.
"The EHG/BedouinB/MA1 model for the Yamnaya doesn't produce any good fits"
I'm probably missing something but it's never made sense to me for Bedouin to be the best marker for first farmers unless there were two sets and the Levant version were later over-run by a second version leaving the Bedouin as a refuge population.
In which case the second set would be the set that was part of Yamnaya.
By far the best qpAdm fit for Yamnaya thus far...
Samara_HG 0.514
Georgian 0.486
The problem is that Georgians have quite a bit of ANE. The question is, where did they get it?
@ Davidksi
"The problem is that Georgians have quite a bit of ANE. The question is, where did they get it?"
...and when ??
Probably northern Iran , during the Kura-Araxes horizon;
Or the north..
Dave how you going in separating the different "types" of ANE ?
I don't think there was much, if any, ANE in northern Iran until the Bronze Age.
Ok
Btw Dave; can you see my question (#6) on your precious admixture post
Can you repeat the question?
Yes Dave,
What are the reference studies you actually used to creat your K15s for the modern populations?
And have you included the new autosomal DNA study for Balkan populations by K Tamberts et al?
Re northern Iran- that is part of central Asia; arguably the "homeland" of R1 -, ANE rich peoples. Whether this remained so after the Holocene drying period is a different question.
@David
''I don't think there was much, if any, ANE in northern Iran until the Bronze Age.''
Yeah You Wish.....
Mike,
Most of my references samples come from here.
http://www.ncbi.nlm.nih.gov/gds
I did use Balkan samples in the making of the K15. They weren't from the paper you site, but that's not important, because the new Balkan samples don't offer any new variation.
OK Thanks
Can you include the Macedonians (you seem to have forgot them).
I am doing some comparisons of modern EE.
@ Alberto
You previously posted a link about the Holocene aridification of central Asia. Do you remember which it was ?
@Mike
You Know about the 5.9,8.2 Etc Kilo Year events right?
http://en.wikipedia.org/wiki/8.2_kiloyear_event
Yeah I do .
But was after specific academic reference for it.
Hey David and Matt,
It looks like the farmers and hunters are too alike to get real mixture with Dstats. There was no mixture from Spain into Hungary before 4500BCE or maybe ever. It is showing admixture solely on Spain having more WHG, from what I can tell. We shall see about the last part, but here you go...
HungaryGamba_EN LBK_EN Spain_EN Chimp -0.0186 -6.545 16209 16825 347882
result: HungaryGamba_EN LBK_EN Spain_MN Chimp -0.0222 -7.517 15902 16624 343551
What do you guys think?
Chad,
You're probably right. But just to make sure everything's ticking over OK, try and repeat some runs from Haak et al. and see if you're getting the same results.
Will do
Mine
LBK_EN Spain_EN Loschbour Chimp -0.0116 -3.015 15817 16188 344777
Theirs
LBK_EN Spain_EN Loschbour Chimp -0.00108 -3.0
David, you need to use HungaryGamba_EN for your Early farmer. It is more Near Eastern than LBK_EN. I'll show you a couple of things, plus playing around with the samples I use.
@ Chad, so I think HungaryGamba_EN LBK_EN Spain_EN Chimp being negative indicates that Spain_EN is more closely related to LBK_EN than it is to HungaryGamba_EN.
You could run the other pairs : HungaryGamba_EN Spain_EN LBK_EN Chimp and Spain_EN LBK_EN HungaryGamba_EN Chimp, to add more detail to their relationships.
Not sure how you'd tell if this was mediated by extra WHG admixture to Spain_EN and/or LBK_EN compared to HungaryGamba_EN though - maybe we could check that with the qpAdm application?
@Mike
I'm not sure it was anything very specific about the aridification process. Just some links to the Kopet Dag/Anau sites that show neolithic settlements since the 7th millennium BCE (as an example of early and successful neolithic development in Central Asia). All make reference to the aridification, but no specific study about it.
This one about farming in the region make some good remarks about climate in the final conclusions:
http://www.persee.fr/web/revues/home/prescript/article/paleo_0153-9345_1994_num_20_2_967
Other links I posted:
http://www.persee.fr/web/revues/home/prescript/article/paleo_0153-9345_2002_num_28_2_4744
http://www.heritageinstitute.com/zoroastrianism/nisa/anau.htm
The event that seems to better coincide with the large migrations could be the 5.9K event:
http://en.wikipedia.org/wiki/5.9_kiloyear_event
@Davidski
"The problem is that Georgians have quite a bit of ANE. The question is, where did they get it?"
One thing we know is that it couldn't be from EHG. There had to be a population with high levels of ANE but without (or low) WHG ancestry somewhere.
My best guess has always been Central Asia. Before the Neolithic they might have been quite pure ANE, but the Neolithic expansion from the Fertile Crescent not only went West (creating EEF in Europe, a mix of NE+WHG), but also East (creating this other population that was a mix of NE+ANE).
The moving from the East Caspian to the West Caspian doesn't seem too difficult, once the conditions in the East Caspian became unsustainable for such population.
With the ancient genomes we have, the ones we lack, and what we know from modern populations, this seems the most reasonable explanation for me.
@ Alberto
Thanks for those refs.
Your proposal certainly seems plausible. What exactly is your definition of 'Central Asia'; and how much do we know about the Mesolithic and Neolithic in central Asia. Is there good evidence of an outmigration from the region ?
Ultimately, aDNA samples from Kazakhstan, Caucasus, Near East, Iran, and India will tell us where this 'West Asian' or 'other ANE' is from.
@Alberto
''The event that seems to better coincide with the large migrations could be the 5.9K event:''
Yes.
@Mike
''What exactly is your definition of 'Central Asia'; and how much do we know about the Mesolithic and Neolithic in central Asia. Is there good evidence of an outmigration from the region ?''
There is the Established migration patterns from the Zarzian-Zagros area to the Urals from Mesolithic to Chalcolithic period it seems, anyway C Asia is a Big area i will dig more but certainly Climate had a basal role in Migrations specially in Neolithic and after....
''Ultimately, aDNA samples from Kazakhstan, Caucasus, Near East, Iran, and India will tell us where this 'West Asian' or 'other ANE' is from.''
Yes we are living in the emerging era of aDNA and we can certainly expect that to happen at least before this decade ends...
Yes, plausible or reasonable is unfortunately the furthest we can go, because unfortunately hard data is mostly missing. No ancient DNA and little archaeology from the region to really have a solid hypothesis.
By "Central Asia" I mostly refer to the region east of the Caspian, though it probably includes neighbouring areas. There seems to be a difference between Turkmenistan/North East Iran (with clear Near Eastern Neolithic influence) and regions further north and east (Uzbekistan, Tajikistan...) where there probably weren't much crops, but at least in Uzbekistan there were domesticated animals. There is not natural border between Uzbekistan and Kazakhstan, so to which extent this "herding" extended north is uncertain to me.
From Afghanistan there seems to be no data at all from these early stages.
The evidence of outmigration is again based on scarce data. The archaeology seems to imply an abandonment of settlements due to climatic changes. But the best evidence should be the places that received the migration. We now have proof that the Steppe received a migration of a population with this genetic profile, and we know that it affected all of Europe. We also know that the people who went to South Asia had this genetic profile.
We lack the proof that Central Asia was actually the origin. But looking at a map doesn't leave many other options.
Hopefully an ancient genome from the region can confirm it at some point (not far away), and that might trigger more interest in finding archaeological data to support it.
Alberto,Mike
For some reason i'm unable to load this, but do tell is this helpful?
http://www.preventionweb.net/files/12033_CCCAdec2009.pdf
@ Alberto
"With the ancient genomes we have, the ones we lack, and what we know from modern populations, this seems the most reasonable explanation for me."
Surely, the ultimate source of this signal, esp. for south of the Caspian and east of the Black Sea could not have been from Yamnaya if Yamnaya is 60 - 70 % 'native EE' - however we name its sub-components.
But its extremely enthralling to know that it appears that groups like Patterson and Kristiansen are already onto it.
Matt,
I've already tested it. Spain en has more WHG than Hungary and LBK. The fact that it allows me to mix a Spaniard at 5100 or 3500BCE, with Germans at 4000-5500BCE, to create a mix mash of Hungarians more than 6500 years old, is enough for me. That mixing event never happened. I'm doing the same thing with swapping in and out samples. It varies based on the WHG.
Nirjhar,
the paper is about current climatic conditions in the region and the human influence on them (Greenhouse Gas emissions, etc..). Historical data only goes back to 1950. In any case, it does underline the still ongoing process of desertification in the area and how it affects the population.
Mike, indeed. A lot of ancient DNA is on the way: Greece, Maykop, Kazakhstan, India, Sumeria... So this year will be quite exciting in this respect.
I can make the Hungarian samples more Near Eastern than LBK by removing a few MN from the test. I'll post a few things.
@Alberto
By "Central Asia" I mostly refer to the region east of the Caspian, though it probably includes neighbouring areas. There seems to be a difference between Turkmenistan/North East Iran (with clear Near Eastern Neolithic influence) and regions further north and east (Uzbekistan, Tajikistan...) where there probably weren't much crops"
turkmenistan has always been an arid place and has never supported huge populations. As for Uzbekistan, its the south east areas bordering Afghanistan, Tajiks where populations are high. Kazakhstan has low population density as well.
The introduction of livestock and farming here is from the south. David has thrown away some interesting signals from his component. One of the few clearly intrusive signals in Yamnaya and europe is the south asian one. The south asian is diluted by Iranian demographic impact, which would probably have a little less ASI, ANE and a bit more wet asian.
HungaryGamba_EN Spain_EN LBK_EN Chimp -0.0146 -4.556 16340 16825 347882
LBK_EN HungaryGamba_EN Spain_EN Chimp 0.0186 6.545 16825 16209 347882
As you can see. It says that people from Germany mixed with people from Hungary to make the Spaniards. Which, again, did not happen. The hunters and farmers are very homogeneous. I will look for specific differences though.
I was also able to create something more Near Eastern than LBK_EN, by mixing Starcevo_EN, LBKT_EN, and KO2 from HungaryGamba_EN. See below.
HungaryGamba_EN1 LBK_EN Loschbour Chimp -0.0158 -2.345 6480 6688 140817
I have lots of other interesting results to show.
HungaryGamba_EN1 LBK_EN Loschbour Chimp -0.0129 -1.765 5579 5724 121704
^^(With Starcevo, KO2)
@Alberto
''the paper is about current climatic conditions ''
:P
@postneo
"turkmenistan has always been an arid place and has never supported huge populations."
But do we know this as a fact? Or just based on the lack of archaeological data? The settlements that have been excavated did seem quite big for the time.
"Kazakhstan has low population density as well."
Yes, Kazakhstan couldn't be the main source of this migration because of this. The same argument goes against the Pontic steppe as the origin.
"One of the few clearly intrusive signals in Yamnaya and europe is the south asian one."
Yes, true. Contacts with South Asia are likely, as with the other neighbouring populations (Near Eastern to the east, EHG to the north). By the data we have, I suggested that this Central Asian population could be in the ballpark of:
40% ANE
40% ENF
10% South Asian
10% WHG
with varying levels depending on the location.
But do you suggest that South Asia could actually be the source of this migration?
MA1 Karelia_HG Loschbour Chimp -0.0658 -7.428 10782 12300 242938
MA1 Karelia_HG Japanese Chimp -0.0100 -1.434 11177 11404 245167
MA1 Karelia_HG Yakut Chimp -0.0143 -2.078 11191 11515 245167
MA1 Karelia_HG Dai Chimp -0.0064 -0.922 11226 11372 245167
MA1 Karelia_HG Han_NChina Chimp -0.0109 -1.544 11163 11410 245167
Karelia_HG Karitiana Japanese Chimp -0.0963 -19.750 14941 18125 341554
Karelia_HG Karitiana Yakut Chimp -0.0751 -15.537 15325 17812 341554
Karelia_HG Karitiana Dai Chimp -0.0889 -17.752 15026 17958 341554
Karelia_HG Karitiana Han_NChina Chimp -0.0922 -18.489 14998 18046 341554
MA1 Karitiana Japanese Chimp -0.1056 -18.602 10888 13458 253297
MA1 Karitiana Yakut Chimp -0.0889 -16.064 11114 13284 253297
MA1 Karitiana Dai Chimp -0.0947 -16.615 11003 13304 253297
MA1 Karitiana Han_NChina Chimp -0.1025 -17.992 10914 13406 253297
SwedenSkoglund_MHG Karitiana Japanese Chimp -0.1326 -11.589 1400 1828 31963
SwedenSkoglund_MHG Karitiana Yakut Chimp -0.1074 -9.423 1445 1792 31963
SwedenSkoglund_MHG Karitiana Dai Chimp -0.1180 -10.222 1415 1794 31963
SwedenSkoglund_MHG Karitiana Han_NChina Chimp -0.1290 -11.143 1403 1819 31963
SwedenSkoglund_NHG Karitiana Japanese Chimp -0.1069 -22.553 14611 18107 325311
SwedenSkoglund_NHG Karitiana Yakut Chimp -0.0906 -19.397 14913 17883 325311
SwedenSkoglund_NHG Karitiana Dai Chimp -0.0998 -20.391 14691 17949 325311
SwedenSkoglund_NHG Karitiana Han_NChina Chimp -0.1030 -21.537 14683 18054 325311
Loschbour Karelia_HG Dai Chimp -0.0131 -2.165 15341 15750 338485
Loschbour Karelia_HG Karitiana Chimp -0.0552 -8.370 14917 16659 338485
Loschbour Karelia_HG Han_NChina Chimp -0.0145 -2.434 15349 15801 338485
Loschbour Karelia_HG Japanese Chimp -0.0145 -2.511 15337 15789 338485
LaBrana1 Loschbour Karelia_HG Chimp -0.0115 -1.426 13350 13660 320734
LaBrana1 Loschbour MA1 Chimp 0.0001 0.006 9826 9826 237767
LaBrana1 Loschbour Dai Chimp 0.0016 0.271 13469 13424 331055
LaBrana1 Loschbour Han_NChina Chimp 0.0004 0.067 13478 13467 331055
LaBrana1 Loschbour Finnish Chimp -0.0181 -3.171 13597 14098 331055
LaBrana1 Loschbour Chuvash Chimp -0.0138 -2.458 13567 13947 331055
LaBrana1 Loschbour Yakut Chimp -0.0018 -0.318 13484 13534 331055
Chad: As you can see. It says that people from Germany mixed with people from Hungary to make the Spaniards. Which, again, did not happen.
I don't think it does, does it?
With:
HungaryGamba_EN Spain_EN ; LBK_EN Chimp -0.0146 -4.556 16340 16825 347882
the fact the stat is negative just means that the second population (Spain_EN) listed is more related to the third (LBK_EN) than the first (HungaryGamba_EN) is.
Doesn't mean that the second population is a mixture of the first and third. That could happen because the two populations form a clade and split off together or perhaps through admixture from an alike source. It's not a test of whether admixture actually happened in any of the populations.
Same with your previous stats:
LBK_EN HungaryGamba_EN Spain_EN Chimp 0.0186 6.545 16825 16209 347882
HungaryGamba_EN LBK_EN Spain_EN Chimp -0.0186 -6.545 16209 16825 347882
which are just simple reverses of one another, just showing again that LBK_EN is more related to Spain_EN than HungaryGamba_EN is.
When you compare the pair:
HungaryGamba_EN LBK_EN Spain_EN Chimp -0.0186 -6.545 16209 16825 347882
HungaryGamba_EN LBK_EN Spain_MN Chimp -0.0222 -7.517 15902 16624 343551
what that means, if I understand correctly, is that LBK_EN is closer to Spain_EN than HungaryGamba_EN, but LBK_EN is even more even more disproportionately close to Spain_MN compared to HungaryGamba_EN.
But why would Spain and Germany be closer other than for WHG? German LBK came from around Hungary. Spanish farmers didn't come from Hungary, as far as we know, but by sea. It should be more about Spain EN having more WHG than HungaryGamba EN and LBK EN. Also, HungaryGamba EN having less WHG than LBK EN.
The only possible scenario I see is some Adriatic Cardium mixing into folks on their way to Germany. I have a European HG structure pretty well completed. I am working on a farmer one now. Would you like to see all of the hunter stats?
@postneo
From a link I posted above:
"there are two points of view about the history of Central Asian climate. According to the first, the modern-day phase of arid and hot climate was preceded by a relatively humid climatic phase named the Lyavlyakan pluvial period, which lasted approximately from the 8th until the 3rd millennium B.C. The second point of view holds that climatic conditions did not alter during the Holocene and were rather arid. However, now we have a quantity of facts indicating that a remarkably more humid climatic phase took place all over the arid zone of the Old World in the middle of the Holocene. Our materials also can serve as indirect evidence of this view. In different parts of Asia, pluvial climatic phases existed during different periods. For the south of Turkmenia humidity occurred around the 6th millennium B.C. From the 5th millennium a process of aridification began again here. The 3rd millennium B.C. was a turning point in the history of the Kopet Dagh foothills. The ancient farming culture achieved the most prosperity here. But beginning from the end of the 3rd millennium B.C., a process of regression caused by further climatic aridification took place. A final decline occurred here in the middle of the 2nd millennium B.C., when the climate became extremely arid."
Chad, I think you're probably right about WHG admixture explaining why Spain_EN and LBK_EN seem more related to either with HungaryGamba_EN, although your other scenario is there as well, whether or not archeology tells us either way I don't know. Perhaps I'm just being pedantic. ;)
I am working on a farmer one now. Would you like to see all of the hunter stats?
Yeah, sure it'd be interesting whenever you're ready with them.
Btw, I thought the stats:
LaBrana1 Loschbour Karelia_HG Chimp -0.0115 -1.426 13350 13660 320734
LaBrana1 Loschbour Finnish Chimp -0.0181 -3.171 13597 14098 331055
LaBrana1 Loschbour Chuvash Chimp -0.0138 -2.458 13567 13947 331055
Seemed new and a little unexpected (maybe on the margins) for showing some possible structure to different WHGs relatedness to EHG and Northeast Europe.
The shifts over the same stats between MHG to NHG to East Asians seem also to show MHG less related to East Asians. I guess SwedenSkoglund_MHG SwedenSkoglund_NHG (Asian) Chimp would confirm that pattern again.
Yes, also maybe more Kostenki in the NHG.
I'll post the link here in just a sec. I'm doing more stuff now.
Here Matt. Go over this. Let me know what you think. Thanks!
https://drive.google.com/?utm_source=en&utm_medium=button&utm_campaign=web&utm_content=gotodrive&usp=gtd<mpl=drive&pli=1#my-drive
Here is something that has me a bit unease about this new K6 calculator.
The percentage of Yamnaya_related ancestry in the following populations:
Basque_French(n=21)=23.57%(min: 19.87% -max: 26.80%)
Central_Greek(n=10)=19.67%(min:13.31%-max:24.56%)
Central_Sicilian(n=2) 11.81%(min:10.99%-max:12.63%)
East_Sicilian(n=11) 12.35% (min:8.15%-max:16.17%)
West_Sicilian(n=18) 14.24% (min:11.32%-max:18.08%)
Now per the West_Eurasian_K8 calculator the percentage of ANE in the same populations:
Basque_French(n=21)=5.95%(min: 4.23% -max: 7.38%)
Central_Greek(n=10)=10.55%(min:8.37%-max:12.04%)
Central_Sicilian(n=2) 7.52%(min:7.43%-max:7.61%)
East_Sicilian(n=11) 7.72% (min:5.85%-max:8.73%)
West_Sicilian(n=18) 8.09% (min:6.68%-max:10.49%)
If we are to assume that the primary vector for ANE in Southern Europe is Yamnaya. How can Basque_French have higher Yamnaya related ancestry than the aforementioned groups? This pattern repeats itself with other groups, I just didn't want to waste to much space in posting it. It's not just Southern Europeans: for example while Scotts have ~3x the amount of ANE per K8 that Basque_French have, they do not have 3x the amount of Yamnaya related ancestry not even 2x.
Because Basques didn't receive the additional flow from West Asia in the Metal Ages, like Italy and the Balkans.
The Near Eastern Cluster on K6 could carry some eastern stuff too, as populations with 8-10% ANE in West Asia have about zero Yamnaya.
Matt,
Here are some more to chew over.
LBK_EN Spain_EN Tunisian Chimp 0.0002 0.069 15670 15664 347888
Spain_EN LBK_EN Tunisian Chimp -0.0002 -0.069 15664 15670 347888
HungaryGamba_EN LBK_EN Tunisian Chimp -0.0082 -3.313 16015 16279 353595
HungaryGamba_EN Spain_EN Tunisian Chimp -0.0085 -2.859 15740 16011 348039
Starcevo_EN Spain_EN Tunisian Chimp 0.0026 0.372 4546 4522 101246
Spain_EN LBK_EN Loschbour Chimp 0.0116 3.015 16188 15817 344777
Spain_EN HungaryGamba_EN Loschbour Chimp 0.0112 2.582 16357 15995 344927
LBK_EN HungaryGamba_EN Loschbour Chimp -0.0013 -0.353 16453 16494 350465
Starcevo_EN LBK_EN Loschbour Chimp -0.0050 -0.607 4638 4685 100720
Starcevo_EN LBK_EN Ju_hoan_North Chimp -0.0064 -0.888 3514 3559 101642
LBK_EN Spain_EN Ju_hoan_North Chimp -0.0007 -0.256 12249 12266 347888
HungaryGamba_EN LBK_EN Ju_hoan_North Chimp -0.0051 -1.982 12608 12737 353595
Starcevo_EN LBK_EN Mbuti Chimp -0.0071 -1.030 3619 3671 101642
LBK_EN Spain_EN Mbuti Chimp 0.0013 0.473 12603 12571 347888
HungaryGamba_EN LBK_EN Mbuti Chimp -0.0086 -3.477 12877 13100 353595
Esperstedt_MN Baalberge_MN Yamnaya Chimp 0.0141 1.855 6419 6241 137066
LBK_EN Baalberge_MN Yamnaya Chimp -0.0051 -1.054 8693 8782 186674
LBK_EN Starcevo_EN Yamnaya Chimp 0.0084 1.163 4702 4623 101573
Spain_EN LBK_EN Yamnaya Chimp 0.0017 0.578 16070 16017 347306
HungaryGamba_EN LBK_EN Yamnaya Chimp -0.0116 -4.315 16300 16682 352535
Corded_Ware_LN Bell_Beaker_LN Spain_EN Chimp -0.0073 -1.905 16303 16542 343948
Corded_Ware_LN Bell_Beaker_LN LBK_EN Chimp -0.0046 -1.303 16581 16733 347638
Corded_Ware_LN Bell_Beaker_LN HungaryGamba_EN Chimp -0.0083 -2.260 16444 16720 347740
Corded_Ware_LN Bell_Beaker_LN Starcevo_EN Chimp -0.0068 -0.976 4799 4865 101163
Esperstedt_MN Baalberge_MN Spain_EN Chimp 0.0185 2.301 6526 6289 136087
Esperstedt_MN Baalberge_MN LBK_EN Chimp 0.0297 3.920 6662 6277 137142
Esperstedt_MN Baalberge_MN SwedenSkoglund_MN Chimp -0.0060 -0.521 4298 4350 91918
Spain_EN LBK_EN SwedenSkoglund_MN Chimp 0.0044 1.095 11333 11233 240801
HungaryGamba_EN HungaryGamba_MN Bell_Beaker_LN Chimp -0.0069 -1.356 12995 13175 277692
HungaryGamba_MN HungaryGamba_LN Bell_Beaker_LN Chimp 0.0114 1.828 10711 10470 225225
Esperstedt_MN HungaryGamba_LN Bell_Beaker_LN Chimp 0.0396 4.950 8673 8012 176303
Baalberge_MN HungaryGamba_LN Bell_Beaker_LN Chimp 0.0205 2.400 5847 5612 120675
LBK_EN HungaryGamba_EN Baalberge_MN Chimp -0.0038 -0.607 7109 7164 150571
HungaryGamba_EN HungaryGamba_MN Baalberge_MN Chimp 0.0079 1.084 7191 7078 150591
HungaryGamba_EN HungaryGamba_LN Baalberge_MN Chimp 0.0147 1.400 4714 4578 98164
HungaryGamba_MN HungaryGamba_LN Baalberge_MN Chimp 0.0054 0.582 5803 5740 121329
HungaryGamba_EN = Starcevo_EN, LBKT, KO2
HungaryGamba_MN = NE1, 2, 3, 4, 5
HungaryGamba_LN = NE7
@Chad: "But why would Spain and Germany be closer other than for WHG?"
That seems a good potential reason. Another possible reason could be admixture of La Hoguette (of possible Cardial origin) into LBK. In any case it seems that the overall data is a bit inconclusive as in general ENE samples tend to form a single cluster.
"The only possible scenario I see is some Adriatic Cardium mixing into folks on their way to Germany".
Archaeologically speaking this doesn't seem to make much sense. The Dinaric range (roughly Bosnia) acted as buffer and was only settled on a second phase, when both LBK and Cardial were already settling the Western areas. More likely (if anything at all) this Cardial-LBK contact took place around the Rhine with the La Hoguette/LBK overlap, way too often ignored (what makes Germany look as "LBK only" when in reality it was largely a mixed area).
The other possibility would be extra WHG admix but other data seems to discard this option before the "Atlantic bounce".
La Hoguette maps online:
1. http://www.archaeologie.bl.ch/Pages/Museum/auston/auston_2_2.jpg (dots are LH findings, shaded areas correspond to LBK and Cardial)
2. http://s27.postimg.org/k1rv6b8yr/Spread_of_Early_Neolithic_Farming_in_Europe_Resize1.jpg (here LH is directly represented as part of "Mediterranean Neolithic" in blue).
@Jean Lohizun: Excellent finding! There's something fishy with this Yamna component and/or with ANE (or both).
@Because "Basques didn't receive the additional flow from West Asia in the Metal Ages, like Italy and the Balkans".
Nobody received that extra-flow you imagine "in the Metal Ages" (except for whatever impact Phoenician colonies and the debated Etruscan migration might have). However the Balcans did receive a second wave from West Asia c. 5000 BCE (Vinca-Dimini, related to Tell Halaf) and they did influence Italy in the Chalcolithic and surely also later.
Anyways, what Jean underlines is that Yamna component (as in this run by Davidski) has not any clear direct correlation with ANE (as per Lazaridis), what indicates that either "Yamna" is hiding other elements (surely) or "ANE" is too slippery to be of any use (almost certain) or both.
PS- If the Vinca-Dimini flow was higher in ANE, as would correspond to to a more Northernly West Asian origin than Thessalian Neolithic (probably Levantine on its West Asian fraction) but had otherwise no direct Yamna/Kurgan relationship... would it work?
PS2- Nah, it doesn't work for Spaniards, who have no apparent Vinca-related influence.
Maju,
I'll see if I can test that out. I'll try to group Early LBK by an Eastern and Western group if possible.
A couple of things I am noticing.
It seems like Loschbour has more of the Kostenki basal
However, KO1 triggers the Neolithic basal more. It seems certain that he has EEF, and there could very well be some in Loschbour and LaBrana, but LaBrana scores just under Loschbour, but not with significant Z score. KO1's Z score is pretty good, something like -2.8, with EEF.
Loschbour seems to pull East Eurasian and other non-EEF, non-Kostenki stuff too.
There looks like an increase in both Kostenki stuff, and EHG in the NHG of Scandinavia. I will keep plugging away.
@Chad: Kostenki, like Mal'ta, are old enough and of diffuse affinity, to trigger nearly anything.
"There looks like an increase in both Kostenki stuff, and EHG in the NHG of Scandinavia".
That's an interesting observation. It would make sense if Pitted Ware, as I understand, is of Dniepr-Don derivation.
LaBrana1 Loschbour BantuSA Chimp 0.0001 0.020 11387 11385 331055
Loschbour HungaryGamba_HG BantuSA Chimp -0.0103 -1.713 8208 8379 241456
LaBrana1 HungaryGamba_HG BantuSA Chimp -0.0097 -1.546 8213 8373 230628
LaBrana1 Loschbour Estonian Chimp -0.0201 -3.571 13616 14175 331055
Loschbour HungaryGamba_HG Estonian Chimp -0.0040 -0.643 10074 10154 241456
LaBrana1 HungaryGamba_HG Estonian Chimp -0.0252 -3.897 9841 10350 230628
LaBrana1 Loschbour Mordovian Chimp -0.0168 -3.018 13603 14069 331055
Loschbour HungaryGamba_HG Mordovian Chimp -0.0044 -0.726 10029 10118 241456
LaBrana1 HungaryGamba_HG Mordovian Chimp -0.0229 -3.623 9830 10290 230628
LaBrana1 Loschbour Iceman Chimp -0.0155 -1.903 13298 13716 329180
Loschbour HungaryGamba_HG Iceman Chimp -0.0067 -0.773 9764 9896 239949
LaBrana1 HungaryGamba_HG Iceman Chimp -0.0255 -2.840 9567 10067 229422
LaBrana1 Loschbour Karelia_HG Chimp -0.0115 -1.426 13350 13660 320734
Loschbour HungaryGamba_HG Karelia_HG Chimp -0.0089 -1.009 9756 9930 233238
LaBrana1 HungaryGamba_HG Karelia_HG Chimp -0.0185 -2.152 9644 10008 223756
LaBrana1 Loschbour Samara_HG Chimp -0.0078 -0.885 8189 8318 195253
Loschbour HungaryGamba_HG Samara_HG Chimp -0.0230 -2.358 5867 6144 142264
LaBrana1 HungaryGamba_HG Samara_HG Chimp -0.0314 -3.060 5896 6278 137150
LaBrana1 Loschbour MA1 Chimp 0.0001 0.006 9826 9826 237767
Loschbour HungaryGamba_HG MA1 Chimp -0.0073 -0.764 7104 7209 173320
LaBrana1 HungaryGamba_HG MA1 Chimp -0.0102 -1.034 7052 7198 166225
LaBrana1 Loschbour Karitiana Chimp -0.0058 -0.863 13503 13662 331055
Loschbour HungaryGamba_HG Karitiana Chimp -0.0056 -0.773 9841 9953 241456
LaBrana1 HungaryGamba_HG Karitiana Chimp -0.0101 -1.313 9809 10009 230628
As you can see, when EEF is involved, it is KO1, slightly ahead of Loschbour.
It could be that LaBrana1 has a touch of Sub-Saharan. He is closer to Loschbour in "SSA" than in "Iceman". Loschbour has a decent Z score over LaBrana in the Iceman, but Loschbour is not selected over LaBrana for BantuSA.
Also, there is a favoritism of Amerindian/EHG type stuff over MA1, to make the difference between LaBrana and Loschbour, but not significant. Maybe LaBrana has almost zero ENA compared to the other two, while having some Sub-Saharan, and lower EEF and Kostenki than the other two. It does look like there could be a cline towards Modern Europeans as you go east and southeast in Mesolithic Europe.
I'll keep looking for recurring themes.
Here is more:
French_HGDP(n=28)
Yamnaya_related ancestry=> Average: 32.67% min: 25.54%-max:40.78%
ANE: Average: => Average: 11.52% min: 7.19%-max:15.43%
ANE/Yamnaya_related=>Average: 0.3526 min: 0.1763-max:0.6041
Polish(n=18)
Yamnaya_related ancestry=>
Average: 47.70% min: 41.90%-max:50.43%
ANE: Average: => Average: 16.81% min: 14.56%-max:18.06%
ANE/Yamnaya_related=>Average: 0.3524 min: 0.2887-max:0.4310
Basque_French(n=21)
Yamnaya_related ancestry=> Average: 23.57% min: 19.87% -max: 26.80%
ANE: Average: => Average: 5.95% min: 4.23% -max: 7.38%
ANE/Yamnaya_related=>Average: 0.2524 min: 0.1578-max:0.3714
@jeanlohizun and Maju
The hypothesis I was trying to see if works is if the Balkans and Italy could have received the ANE from the "other" population that was part of Yamnaya. The one that David found a best match with Georgians.
Maju, let's say that Maykop (and even more likely Kura Araxes) were a Georgian like population (that is, the other half of Yamnaya that was not EHG).
Couldn't this population have gone to the Balkans through Anatolia? For example, Cernavoda (or later Ezero) could come from there without going around the north coast of the Black Sea. Genetically that could explain why SE Europeans have more ANE but less Yamnaya (or actually no Yamnaya). Archaeologically I don't know (if they moved through Anatolia, they didn't leave traces?). Any opinion about this possibility?
jeanlohizun,
Thanks for the stats.
Please keep in mind that the Yamnaya component is not ANE, and ANE is not the Yamnaya component.
For instance, in my tests Pashtuns get around the same levels of Yamnaya and ANE, and this is probably because they received a good portion of their ANE from groups not closely related to the Yamnaya.
Some structure within EHG. Nothing terribly significant, but interesting, none the less.
It seems that when it is straight ENA, there is no favoritism. When it is ENA and "ANE", Karelia is very minutely favored, but nothing significant. However, when we use West Asian and South Central Asians, Samara is well ahead, but not quite significant.
Karelia_HG Samara_HG Kostenki14 Chimp -0.0041 -0.461 8320 8389 192347
Karelia_HG Samara_HG Karitiana Chimp 0.0061 0.824 9039 8928 203451
Karelia_HG Samara_HG Loschbour Chimp -0.0026 -0.282 9004 9051 201624
Karelia_HG Samara_HG Sindhi Chimp -0.0084 -1.359 8859 9009 203451
Karelia_HG Samara_HG Japanese Chimp 0.0002 0.028 8834 8831 203451
Karelia_HG Samara_HG Han_NChina Chimp -0.0010 -0.144 8834 8852 203451
Karelia_HG Samara_HG Brahui Chimp -0.0089 -1.434 8848 9007 203451
Karelia_HG Samara_HG Pathan Chimp -0.0052 -0.820 8899 8991 203451
Karelia_HG Samara_HG Kalash Chimp -0.0049 -0.743 8933 9020 203451
Karelia_HG Samara_HG Balochi Chimp -0.0089 -1.450 8834 8992 203451
Karelia_HG Samara_HG Lezgin Chimp -0.0083 -1.280 8926 9075 203451
Karelia_HG Samara_HG Georgian Chimp -0.0063 -0.966 8926 9038 203451
Karelia_HG Samara_HG Armenian Chimp -0.0092 -1.404 8880 9045 203451
Karelia_HG Samara_HG Iraqi_Jew Chimp -0.0113 -1.723 8818 9019 203451
Karelia_HG Samara_HG Tajik_Pomiri Chimp -0.0073 -1.120 8919 9050 203451
Karelia_HG Samara_HG Turkmen Chimp -0.0049 -0.763 8891 8979 203451
It does look like MA1 has Kostenki ancestry, but not as much as Loschbour and KO1. Although I do wonder if there is some ENA in MA1, but not Kostenki causing this. The only thing that may dispute that is the fact that Karitiana natives are closer to MA1 in Kostenki than Japanese, by a descent amount.
MA1 Karitiana Kostenki14 Chimp 0.0328 4.389 11636 10896 238432
MA1 Japanese Kostenki14 Chimp 0.0499 7.411 12494 11306 238432
MA1 Yakut Kostenki14 Chimp 0.0437 6.686 12270 11243 238432
MA1 Uzbek Kostenki14 Chimp 0.0357 5.573 12166 11327 238432
MA1 Ust_Ishim Kostenki14 Chimp 0.0571 6.463 12828 11441 237947
MA1 Karelia_HG Kostenki14 Chimp -0.0047 -0.533 10602 10703 231217
MA1 Samara_HG Kostenki14 Chimp -0.0009 -0.083 6425 6436 141063
MA1 Karelia_HG Ust_Ishim Chimp 0.0088 1.029 11150 10954 244665
MA1 Samara_HG Ust_Ishim Chimp 0.0041 0.411 6705 6650 148789
Loschbour MA1 Kostenki14 Chimp 0.0276 3.329 11760 11127 236258
LaBrana1 MA1 Kostenki14 Chimp 0.0246 2.765 11185 10649 226460
HungaryGamba_HG MA1 Kostenki14 Chimp 0.0235 2.354 8117 7744 164971
SwedenSkoglund_MHG MA1 Kostenki14 Chimp -0.0182 -0.825 1051 1090 21983
SwedenSkoglund_NHG MA1 Kostenki14 Chimp 0.0030 0.355 10675 10610 219561
"For instance, in my tests Pashtuns get around the same levels of Yamnaya and ANE, and this is probably because they received a good portion of their ANE from groups not closely related to the Yamnaya."
Do you suppose a big difference in K6_Yamnaya to K8_ANE ratio implies some actual ancestry from the steppe?
Chad
Don't we already know this?
Haak shows that Kostenki is closer to WHG than to samara and Karelia, which makes sense given that it is Hg C; which probably radiated through balkans to west and east europe .
On the other hand; it Ma1 is clearly eastern, and later EHG obviously derive from later movements into EE from central Asia .
@ Alberto
Of course it's possible . There are clear signs of similarities across the Pontic ; from Ikziterpe in northern Turkey to Troy in the west, Cernadova on western Black Sea coast and other sites in Thrace, etc. It is I only assumed that these represent the "Kurganized" defensive forts of the Bronze Age- a wrong assumption at that.
As for whether the Balkans got ANE from anatolia directly ; it Seems unlikely though; unless it was a later movement of M269 carriers who moved west from Central Asia. But now I'm in speculation territory
Gill,
Yes, I think so, otherwise we'd see much higher Yamnaya among South Central Asians, inflated by the high ANE among them, but we don't. There's a nice gradient there from the Volga-Ural to the Pamirs and into the Hindu Kush and Iran. It just makes sense.
Mike,
Jean is comparing apples to oranges.
http://m.genome.cshlp.org/content/early/2015/03/13/gr.186684.114.abstract
"... if the Balkans and Italy could have received the ANE from the "other" population that was part of Yamnaya. The one that David found a best match with Georgians".
Sure. Northern West Asians (i.e. Zagros-Anatolia-Caucasus) seem to have got since long ago more ANE than Southern West Asians (i.e. Palestine and surroundings), much as they do now. The Vinca-Dimini intrusion and other Anatolian-related flows like the likely Tyrsenian one into Italy could well be source of that Highland West Asian element, still West Asian but with more ANE.
It should not have significantly affected Iberia or other parts of Western Europe however. The seemingly tiny Phoenician/Jewish/Arab input in that region should if anything be more Palestinian-like, while historical Greek presence was trivial (Marseilles and a few outposts) and prehistorical one was also surely very small (Mycenaean cultural influence in proto-Iberians did not alter the language nor anything else in any major way, only burial rituals can be identified as an influence and it is bidirectional).
@Alberto (previous comment also for him):
"Couldn't this population have gone to the Balkans through Anatolia?"
I don't see even the slightest hint that could support such scenario. Cernavoda, Ezero, etc. all seem to drink from steppe and local Balcan sources directly, while the expansion of Anatolian IE pops. westwards (in Anatolia only) is tightly related to the formation and growth of the Hittite Empire.
As I said above, the most likely source for what you say is the second Neolithic wave of Dimini-Vinca (Halaf-related), which is much older. As for Italy, probably a combination of Balcanic and Anatolian (Etruscans particularly) waves or influences should be sought for. The Balcanic influences can be tracked to the Chalcolithic (and overlap with influences from the West), continuing later in the South with Greek colonization (not just classical but surely also in Mycenaean times to some extent). These influences appear to be much stronger in Southern and Central Italy than in the North.
@David: "Please keep in mind that the Yamnaya component is not ANE, and ANE is not the Yamnaya component".
Obviously. But shouldn't they in Europe be very strictly correlated? If not (as seems to be the case), then most likely there are other sources of the Yamnaya-like component which are lower in ANE, i.e. there are hidden confounding factors at play.
@Chad, on the new study: notice that they date OoA to ~50 Ka BP which is dramatically wrong (should be ~100 Ka BP and there's no way around it), so their purported Y-DNA bottleneck c. 10 Ka BP is most likely in fact c. 20 Ka BP, i.e. just after the LGM.
They don't have OoA at 50kya. Read it again and look at the tree. Looks like a UP bottleneck or something.
Maju,
I'd say some ANE also entered Europe from west Asia during the Iron Age and middle ages, and was very likely accompanied by a very low ratio of EHG and a high ratio of Near Eastern ancestry.
Maju, you can see in Davidski's PCAs southern and western Spanish and Portuguese cluster right by North Italians. East Med and African components in K15 gradually rise in Iberia as you go east and west. Both those suggest significant west Asian and North African influence not in other parts of Iberia.
@ Maju
"don't see even the slightest hint that could support such scenario. Cernavoda, Ezero, etc. all seem to drink from steppe and local Balcan sources directly,"
Don't confuse what you aren't aware of with what doesn't exist. As I've pointed out above that there Clearly are parallels across the Pontic including anatolia; and to reduce the Pontic forts to steppe derivations is plain wrong.
"Expansion of IE in anatolia linked to Hittites "
Wow . youre so wrong it worries me
There were several IE languages in anatolia which have little to do with Hittite and the expansion of their empire. In fact, the Anatolian IE languages are rather diverse that Ivanov has questioned their immediate genetic relatedness. Clearly IE has a complex (deep) history in Anatolia
Post a Comment